Research Article, J Mar Biol Oceanogr Vol: 8 Issue: 1
Bio-Corrosion, Sulfate-Reducing Bacteria in the Yucatan Peninsula
Arena-Ortiz Ma Leticia*, Mahendhiran Muhilan, Reyes-Sosa Mariela and Ortiz-Alcantara Joanna
Unidad Multidisciplinaria de Docencia e Investigación, Facultad de Ciencias, Universidad Nacional Autónoma de México, Puerto de abrigo s/n, Sisal, Yucatán, México
*Corresponding Author : Arena-Ortiz Ma Leticia
Profesor Titular B, TC, Laboratorio de Estudios Ecogenómicos, UNAM, Parque Científico Tecnológico de Yucatán, Carretera Sierra Papacal-Chuburná, km 5.3 C.P. 97302, Yucatán, México
Tel: (999) 3410867
E-mail: leticia.arena@ciencias.unam.mx
Received: January 19, 2019 Accepted: March 14, 2019 Published: March 20, 2019
Citation: Leticia AM, Muhilan M, Mariela R, Joanna O (2019) Bio-Corrosion, Sulfate-Reducing Bacteria in the Yucatan Peninsula. J Mar Biol Oceanogr 8:1. doi: 10.4172/2324-8661.1000202
Abstract
Microorganisms colonize the engineering materials and could damage them eventually. Sulfate reducing bacteria (SRB, hereafter) are responsible for corrosion in various metals such as carbon steel, stainless steel, iron and some alloys. Biocorrosion causes huge loss every year which causes various socio and economic complications. In anaerobic environments, SRB are active and thus microbiologically influenced corrosion (MIC) could occur. However, molecular mechanisms of MIC are less understood. Therefore, it is important to recognize the microorganisms at the molecular level and their mechanisms associated to biocorrosion, in order to reduce/prevent damages caused by these organisms. Also, the identification of various genera and species related to biocorrosion is needed. In this article, we determined the presence of SRB related to biocorrosion under anoxic conditions in the Yucatan Peninsula of Mexico. Water and sediment samples were collected from a total of twenty-one sampling sites coming under four environmental types (Freshwater upwelling’s, Lagoon, Sea, Sea (Beach) and Wetlands) in Sisal Coastal region of Yucatan State in Mexico. Physicochemical parameters such as pH, salinity, dissolved oxygen, conductivity; total dissolved solids, redox potential and temperature were monitored in situ. 16S ribosomal RNA (rRNA) gene-based sequencing analysis showed that, out of 37 bacterial genera in SRB group, mainly eight anaerobic bacteria were present in both water and sediment samples including Desulfatibacillum, Desulfatitalea, Desulfobacula, Desulfobulbus, Desulfotignum, Desulfotomaculum, Desulfovibrio and Sulfurospirillum. Principal Component Analysis (PCA) showed that at least six variables were correlated between the selected sampling sites. Hierarchical Cluster Analysis (HCA) indicated a minimum of five groups of environmental variables, including outgroup and two major clusters. This is the first study of the presence of sulfate reducing bacteria which could cause biocorrosion in the Yucatan Peninsula of Mexico.
Keywords: 16S ribosomal RNA (16S rRNA) gene sequencing; Sulfate-reducing bacteria (SRB), Anaerobe; Microbiologically influenced corrosion (MIC); Yucatan Peninsula; Mexico
Introduction
Corrosion is a chemical action or oxidation which could produce deterioration of the iron and steel. Anaerobic microorganisms endorse iron corrosion, consuming hydrogen as an electron donor from oil facilities [1]. Iron is usually unstable and easily affected by corrosion which could cause severe damages. Corrosion damages and protection measures result in losses of US $ 4 trillion per year all over the world, and also cause various socio-economic complications and human health problems [2-4]. In the Gulf of Mexico, most of the cases of major pipeline failures are caused by corrosion [5]. In industrial sectors, the inferred costs of metal corrosion affect the Gross domestic product (GDP) value of each country from 4% in developing/industrialized countries and 20% of total cost is because of microbial corrosion mechanisms [6,7].
Microbiologically influenced corrosion (MIC) is a serious threat to essential infrastructure in the steel and pipeline industries. In energy industries, microbiologically influenced corrosion in oil and gas pipelines is responsible for more than 20% of the total corrosion expense which could costs billions of dollars loss every year [8-10]. Anaerobic iron corrosion is one of the major devastating damages on infrastructure, such as oil and gas structures in pipeline systems, refineries and storage terminals [9,11,12]. The bacteria which reduce sulfate and iron are called sulfate reducing and iron reducing bacteria respectively [13]. The biocorrosion caused by Sulfate-reducing bacteria (SRB) has been studied over many years. Formation of iron sulfides (FeS) which could lead to iron corrosion by SRB might reduce corrosion in some cases, whereas increase in others [14,15]. Microbes grow on surfaces of various metals and accomplish various metabolic reactions which promote the deterioration of the substrate. Sulfate-reducing bacteria (SRB) causes biocorrosion under anoxic conditions and the activity of these bacteria in some cases creates serious environmental problems [14,16]. However, their use under carefully controlled conditions could be beneficial, such as purposes both in domestic and industrial and saline wastewater treatment [17,18].
Sulfate reducing bacteria (SRB) are a ubiquitous group of anaerobic bacteria found in sulfate rich environments such as marine, estuarine, sewage, freshwater wetlands, humans and animals [19-21]. SRB are involved in geochemical carbon and sulfur cycles both in land and sea [22-24] and contribute to around 50% of organic matter mineralization in marine sediments [25,26]. These anaerobic bacteria also live in intestinal guts of various animals and humans causing severe damages [27,28]. In anerobic environments SRB performs dissimilatory sulfate reduction in order to obtain energy [29-31]. SRB release Hydrogen sulfide (H2S) as the final product, after the reduction process of sulfate (electron acceptor) and utilize it for anaerobic respiration [32-34]; this H2S is toxic, flammable, corrosive and also contributes to corrosion. Moreover, H2S gas is a serious threat to human beings, transportation pipelines and production places [35] and damages the metals by increasing the iron sulfide concentration which could lead to an increase in the corrosion rate as well. SRB could utilize various organic electron donors for sulfate reduction; for example, Desulfoluna butyratoxydans, sulfate-reducing bacterial strain MSL71T consume formate, butyrate, pyruvate, lactate, malate, ethanol, propanol, butanol, glycerol and H2 [36]. SRB are also involved in anaerobic degradation of crude oil in sulfate-containing environments [37].
Sulfate-reducing bacteria are classified into five phylogenetic lineages, which consist of 220 species of 60 genera [38]. i) Desulfobulbus (all delta subgroup) and the genera Desulfovibrio, Desulfobacterium, Desulfobacter with mesophic proteobacterias [39], ii) thermophilic G-ve bacteria with the genus Thermodesulfovibrio, iii) G+ve Peptococcaceae with the genus Desulfotomaculum, iv) a genus of Archaea named Archaeoglobus (22) and v) finally thermophilic autotroph Thermodesulfobium narugense of the family Thermodesulfobiaceae [40].
The objective of this study was to identify the presence of sulfate-reducing bacteria (SRB) related to biocorrosion in different environmental samples in the Yucatan peninsula of Mexico.
Materials and Methods
Sampling sites
Water and sediment samples were collected from different sites (Freshwater upwellings, Lagoon, Sea, Sea (Beach) and Wetlands) at a total of twenty-one sites near the Sisal Port of Yucatan state in Mexico from March to May and in August 2016 (Table 1). Sampling points were geolocated using a GPS navigation device (Garmin, Olathe, KS, USA). Various physicochemical parameters were used to assess the water quality. Physicochemical data in water including pH, salinity, dissolved oxygen, conductivity, total dissolved solids, redox potential and temperature were collected in situ using a multiparameter probe (YSI, OH, USA, Table 1). Sediment samples were collected in duplicates (20 to 25 cm in depth) using a polypropylene corer of 18 mm diameter and one meter long. The samples were stored in sterile 50 mL conical tubes. Water samples were collected in sterile plastic bottles (3 to 5 L total per site). Samples were transported to the laboratory in an ice box. Pulled column water samples for each sampling site were filtered using a nitrocellulose membrane of 0.45 µm pore size (Millipore, Darmstadt, Germany). Sediment samples and water filters were stored at -20°C until DNA extraction process.
| No | Environment | Sample type (Water/Sediment) | Location (GPS) North/West |
|---|---|---|---|
| 1 | Freshwater upwellings | Water | 21º13'8.2554'/89º53'45.0954'' |
| 2 | Lagoon | Water and sediment | 21º13'35''/89º53'44'' |
| 3 | Lagoon | Water and sediment | 21º13'15''/89º52'58'' |
| 4 | Lagoon | Water and sediment | 21º13'08''/89º52'54'' |
| 5 | Wetlands | Water and sediment | 21º09'33.7''/90º02'51.4'' |
| 6 | Wetlands | Water and sediment | 21º09'30.4''/90º02'42.4'' |
| 7 | Wetlands | Water and sediment | 21º09'46.8''/90º01'52.2'' |
| 8 | Wetlands | Water and sediment | 21º09'53''/90º01'28.5'' |
| 9 (i) | Sea (Beach) | Water | 21º10'03.8''/90º01'54.3'' |
| 9 (ii) | Sea (Beach) | Sediment | 21º10'03.8''/90º01'55.9'' |
| 10 | Sea (Beach) | Water and sediment | 21º10'0.5''/90º02'1.8'' |
| 11 | Sea (Beach) | Water and sediment | 21º09'59.7''/90º02'04.6'' |
| 12 | Sea (Beach) | Water and sediment | 21º09'59.9''/90º02'14.3'' |
| 13 | Sea (Beach) | Water and sediment | 21º09'57.9''/90º02'18.0'' |
| 14 | Sea (Beach) | Water and sediment | 21º09'57.8''/90º02'31.4'' |
| 15 | Sea (Beach) | Water and sediment | 21º09'57.7''/90º02'55.9'' |
| 16 | Sea | Water and sediment | 21º09'34.14''/90º02'53.92'' |
| 17 | Freshwater upwellings | Water and sediment | 21º11'24.7''/89º57'09.2'' |
| 18 | Freshwater upwellings | Water and sediment | 21º11'52.0''/89º56'48.0'' |
| 19 | Wetlands | Water | 21º10'02.1''/90º00'47.8'' |
| 20 | Wetlands | Water | 21º11'54.2''/89º57'09.2'' |
| 21 | Sea | Water and sediment | 21º20'08.36''/90º08'08.61'' |
Table 1: Samples and sampling sites used in this study.
DNA extraction and sequencing
Total DNA purification was performed with previously reported silica adsorption-based method [41]. DNA was confirmed in a 1% agarose gel, concentration and purity were evaluated using both NanoDrop 2000 (Thermo Scientific, Waltham, MA, USA) and Quantus Fluorometer and the Quantifluor dsDNA system (Promega Corporation, Madison WI, USA). 16S rRNA gene sequencing was performed in the Laboratorio Nacional de Genómica para la Biodiversidad (LANGEBIO, Irapuato, Gto., Mexico). Libraries were prepared using the Nextera XT kit (Illumina, San Diego, CA, USA) and sequenced in a MiSeq instrument (Illumina, San Diego, CA, USA) generating 2 × 300 paired-end reads.
Sequence data analysis
Paired reads were merged using PEAR v1.10.5 [42]. Merged reads were filtered using PRINSEQ-lite v0.20.4 [43], with Q20 as quality filter. Primers and adapters were removed using TagCleaner v0.16 [44]. FastQC v0.11.5 (Babraham Institute, UK) was used for quality assessment of raw reads and after the TagCleaner process. One Codex [45] was used for taxonomic assignment. A search to identify bacteria associated to biocorrosion according to previous reports was performed.
Statistical analyses
The physicochemical parameters obtained from environmental sampling sites were analyzed by Primer 6. The environmental variable data were uploaded and then transformed overall by square-root method. Subsequently, Principal Component Analysis (PCA) was performed from the databases of each site to find the correlation between environmental variables. Based on the variation percentage, the most sensible six variables were selected and multivariate analysis was done [46].
In order to find similarities between the samples, we constructed a resemblance matrix with the above-mentioned environmental variables using Primer 6. Subsequently, Hierarchical Cluster Analysis (HCA) of major components was performed [47].
Results
Microbiologically-influenced corrosion (MIC) or biocorrosion causes severe damages in oil and pipeline industries; the causal agent of this problem are Sulfate reducing bacteria [SRB; 14,48,49]. The production of sulfide is highly reactive, corrosive, toxic and causes environmental and economic impact [32,50,51]. In the present study SRB were widely found in both water and sediment samples in the Yucatan peninsula of Mexico. Physicochemical parameters measured in water are presented in Table 2.
| No | Environment | Oxygen (%) | Oxygen concentration (mg/L) | pH | Redox potential | Temperature (°C) | Salinity | Conductivity (mS/m) |
Total dissolved solids (g/L) |
|---|---|---|---|---|---|---|---|---|---|
| 1 | Freshwater upwellings | 4.7 | 0.35 | 7.41 | 77.3 | 26.64 | 1.76 | 3.47 | 2.19 |
| 2 | Lagoon | 86 | 5.25 | 8.67 | 69.9 | 31.10 | 36.03 | 61.07 | 35.55 |
| 3 | Lagoon | 109.4 | 6.4 | 8.83 | 33.9 | 33.3 | 38.42 | 67.22 | 37.72 |
| 4 | Lagoon | 136.5 | 7.78 | 9.02 | -30.1 | 35.04 | 38.40 | 63.22 | 37.75 |
| 5 | Wetlands | 27.3 | 1.78 | 8.08 | 117.5 | 30.01 | 25.82 | 44.88 | 26.39 |
| 6 | Wetlands | 16.7 | 1.16 | 8.58 | -36.2 | 26.91 | 25.76 | 38.92 | 24.41 |
| 7 | Wetlands | 51.5 | 3.61 | 8.71 | -2.6 | 78.33 | 19.02 | 32.69 | 19.99 |
| 8 | Wetlands | 72.9 | 4.93 | 8.57 | -94.5 | 30.32 | 19.72 | 35.02 | 20.69 |
| 9 (i) | Sea (Beach) | 108 | 6.9 | 8.2 | 41.7 | 29.30 | 32.92 | 54.54 | 32.76 |
| 9 (ii) | Sea (Beach) | 106 | 6.8 | 8.23 | 37.8 | 29.30 | 32.85 | 54.23 | 32.69 |
| 10 | Sea (Beach) | 95.8 | 5.87 | 8.46 | 63.9 | 30.59 | 36.57 | 61.35 | 36.01 |
| 11 | Sea (Beach) | 95.1 | 5.83 | 8.86 | 50.8 | 30.61 | 36.64 | 61.49 | 36.08 |
| 12 | Sea (Beach) | 142 | 9.2 | 8.16 | No data | 28.7 | 33.88 | 53.81 | 32.59 |
| 13 | Sea (Beach) | 98.8 | 6.0 | 8.57 | 72.5 | 30.65 | 36.59 | 61.41 | 36.06 |
| 14 | Sea (Beach) | 127 | 8.2 | 8.17 | No data | 28.7 | 32.98 | 54.00 | 32.82 |
| 15 | Sea (Beach) | 120 | 7.8 | 8.07 | 4.01 | 27.9 | 33.01 | 53.23 | 32.82 |
| 16 | Sea | 76.0 | 4.85 | 8.06 | -25.8 | 29.83 | 32.46 | 54.20 | 32.35 |
| 17 | Freshwater upwellings | 29.0 | 2.2 | 6.99 | 85.3 | 25.2 | 2.22 | 34.47 | 2.22 |
| 18 | Freshwater upwellings | 4.0 | 0.3 | 6.90 | 67.0 | 25.0 | 25.0 | 63.57 | 4.13 |
| 19 | Wetlands | 44 | 3.0 | 8.12 | 166.5 | 30.2 | 16.51 | 29.80 | 17.62 |
| 20 | Wetlands | 107 | 6.1 | 8.53 | 100.6 | 33.3 | 36.10 | 63.52 | 35.68 |
| 21 | Sea | 109 | 8.6 | 8.02 | 35.5 | 26.1 | 32.57 | 50.88 | 32.37 |
Table 2: Physicochemical properties of water in the sampling sites.
Generally, SRB grow optimally in the pH range of 5.5-8.5 and inhabit temperatures that vary from 0 to 100°C, with optimum temperature of 24-42°C [38,52]. Although, some SRB like Desulfonatronovibrio hydrogenovorans and Desulfonatronospira thiodismutans can survive at pH>9.5 and pH<5 respectively [33,53,54]. However, at the time of sampling, the pH and temperature ranged from 6.90 to 9.02 and 25.0 to 35.04°C respectively, except in one sample from wetlands (water sample 7) where the temperature of water was 78.33°C, this place is near to the pipeline of the gas station of Sisal, Yucatan (Table 2).
In addition, other parameters such as oxygen percentage, oxygen concentration (mg/L), Redox potential, salt concentration, conductivity (mS/m) and total dissolved solids (g/L) were shown (Table 2). Percentage of oxygen and oxygen concentration (mg/L) varied in different environments; from very low to high concentrations including Freshwater upwellings (0.3 mg/L) to lagoon (7.78 mg/L) and Sea (Beach; 9.2 mg/L). Salinity, conductivity and total dissolved solids were also presented (Table 2).
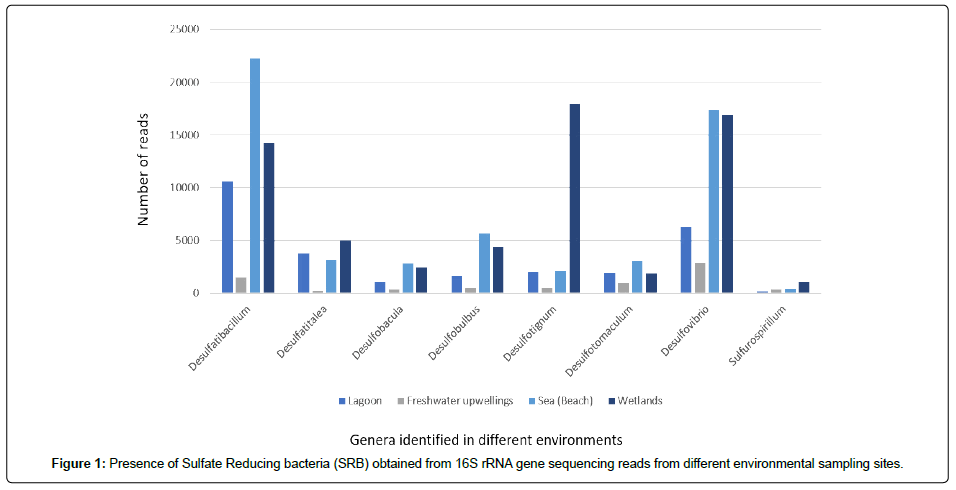
Among the five phylogenetic lineages of SRB [38,55], our 16S rRNA gene sequencing analysis showed that eight genera were identified in twenty-one sampling sites of environments such as Lagoon, Freshwater upwellings, Sea and Wetlands in both water and sediment samples; such as Desulfatibacillum, Desulfatitalea, Desulfobacula, Desulfobulbus, Desulfotignum, Desulfotomaculum, Desulfovibrio and Sulfurospirillum (Figure 1).
Each environment showed different abundance of reads for each genus. Desulfatibacillum reads were the most abundant, in lagoon and sea (Figure 1). Desulfovibrio reads were the most abundant in freshwater upwellings. In wetlands, the most abundant genera in sequencing reads were Desulfotignum, Desulfovibrio and Desulfatibacilum. Sulfurospirillum was the least abundant genus, in all the environments followed by Desulfobacula and Desulfotomaculum.
Desulfotomaculum spp are anaerobic, gram-positive, widely present, heat-resistant spore producing and very diverse in both phylogenetically and physiologically SRB [56-58]. Desulfotomaculum genus was widely found in our sampling sites, except lagoon (2w), wetlands (19w), wetlands (20w) and sea (21w). This genus is composed of 30 species and one subspecies [59]. In our samples, we have found Desulfotomaculum nigrificans species in just one sampling site (3w lagoon); in sediment samples it was found in two places in wetlands (5s) and sea (21s, data not shown).
Desulfovibrio sp. is a genus of SRB, gram negative, anaerobic bacteria, which create MIC in steel pipeline industries [9]. We have found Desulfovibrio desulfuricans ND132 in most of the sampling sites. D. desulfuricans species could be detected in soil, fresh water and salt marshes, environment, and human intestinal flora [60,61]. In our study, Desulfovibrio genus and its various species were found widely when compared with other genera. Desulfovibrio alaskensis was found both in the water and sediment samples from many sites (data not shown). D. alaskensis strains are related to petroleum industries [62], this strain might be found in contaminated petroleum regions through the Gulf of Mexico. Similarly, D. alaskensis G20 bacteria was isolated from a corrosion site of an oil well in Ventura Country, California [35].
In all twenty-one water sampling sites we have found Thermodesulfobium narugense DSM 14796 (Thermodesulfobium genus, data not shown). This T. narugense is an anerobic, acidophilic and thermophilic autotroph SRB species and classified under Firmicutes phylum [40,63], present in the water samples of Freshwater upwellings and in Lagoon. Species: T. narugense DSM 14796 is a strain type SRB that was previously isolated from Narugo hot spring of Miyagi, Japan (40). Whereas in sediment samples, Desulfurobacterium sp. TC5-1 was found only in Lagoon (2s). Desulfobulbus sp. Tol-SR is a species found in all the locations in sediments and in water samples as well (sea 21w, s).
A little variation has been found in water and sediment samples from the same sampling sites. Even at high temperature as 78.33°C, we have found all the eight genera of sulfate reducing bacteria in both water and sediment samples at one place, (Wetlands 7w, s; Table 2). Out of twenty-one sampling sites, we were not able to obtain any data sequences from three sampling sites, such as freshwater upwellings (1s), wetlands (19s, 20s) in sediment samples. We could not obtain amplicons for the above-mentioned sediment samples, because they had not enough concentration of DNA. Therefore, we analyzed only 21 water samples and 18 sediment samples from distinct environments.
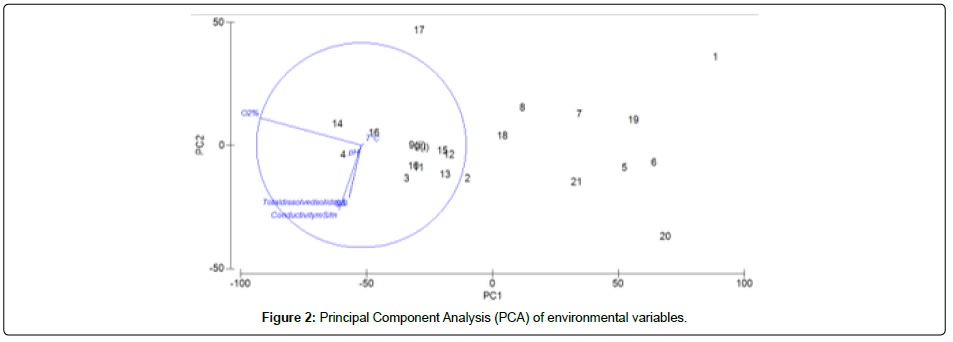
Principal Component Analysis (PCA) from the physicochemical parameters of environmental data sets. Out of eight variables, six were significative and thus considered as principal components, such as oxygen percentage, pH, Temperature, Salinity, Conductivity and Total dissolved solids (Figure 2 and Table 3). Percent variance (% var) and cumulative percent variance (Cum. %) contribution has been showed for each variable. However, the variable Total Dissolved Solids (TDS) g/L did not contribute to the percent variance in this statistical analysis. As indicated in Table 3, oxygen percentage was the principal component among all the variables with the highest percent variance (80.5%) followed by other components.
| No. | Principal Component (PC) | Eigenvalues | % variation | Cum % variation |
|---|---|---|---|---|
| 1 | Oxygen (O2) % | 1.96E3 | 80.5 | 80.5 |
| 2 | pH | 311 | 12.8 | 93.3 |
| 3 | Temperature (T) °C | 117 | 4.8 | 98.1 |
| 4 | Salinity (ppm) | 44.2 | 1.8 | 99.9 |
| 5 | Conductivity (mS/m) | 2.82 | 0.1 | 100.0 |
Table 3: Principal Component Analysis (PCA) of variables from environmental parameters.
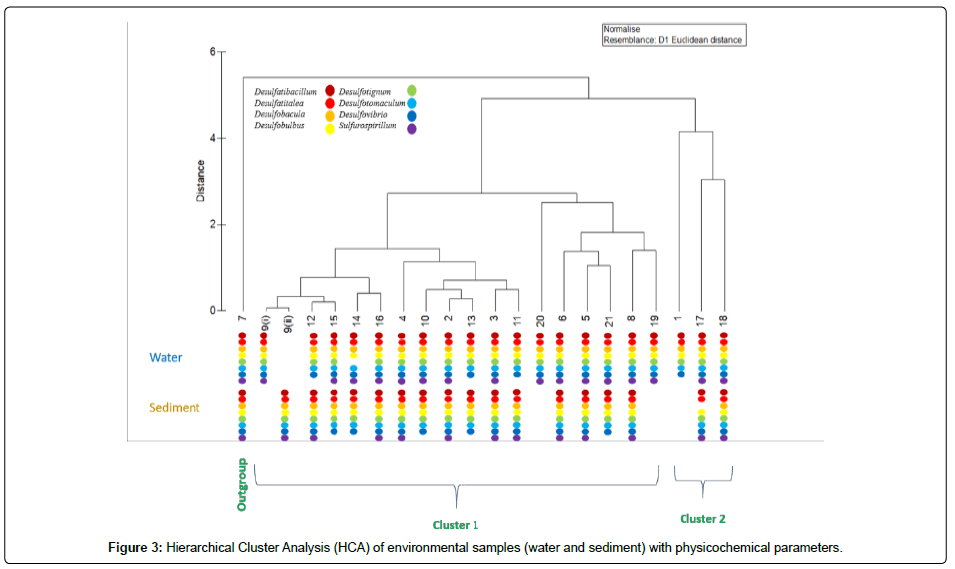
To complement PCA, a Hierarchical Clustering Analysis (HCA) was carried out (Figure 3) in search of a stricter division of groups, establishing similarities between environmental variables and thus facilitating the processing of information. HCA showed in Figure 3 that an out group and two major clusters, such as cluster 1 and cluster 2 were obtained. Out group sample (7 wetlands) was collected near to a gas station were the water level was very low. Thus, the temperature was the highest among all the sampling sites (78.33°C).
Cluster 1 contained three sub-clusters such as sub-clusters 1, 2 and 3. Sub-cluster 1 contained beach and sea samples which had saline water. Sub-cluster 2 included lagoon and beach (sea) samples where salinity was higher. In, sub-cluster 3 most of the samples were from wetlands and just one from sea [21]. In wetlands, the salinity of the water was reduced due to rain fall and thus formed a separate sub-cluster.
Cluster 2 included samples 1, 17 and 18 from freshwater upwelling, where pH values and temperature were relatively low in comparison with other samples. The vegetation was abundant and thus reduced the heat in the ambient. In all the eight SRB genera, Sulfurospirillum was not found in some of the water [1,11-13] and sediment samples [10,14,15,21]. It is interesting that even at high temperature (78.33°C) all the above-mentioned genera were found in both water and sediment samples. Overall, cluster 1 and cluster 2 contained salt water and freshwater respectively (Figure 3).
Thus, statistical analyses such as Principal Component Analysis (PCA) and Hierarchical Cluster Analysis (HCA) showed that samples could be grouped according to the physicochemical parameters measured in water. Eight SRB genera were found in most of the environmental water and sediment samples in all groups.
Discussion
Sulfate-reducing bacteria (SRB) have been considered as typical and ubiquitous in anaerobic environments and one of the most ancient prokaryotes [1,16,51,64]. In tropical mangrove, marine water and sediment samples the temperature is one of the key elements for the healthy microbial niche, metabolic activities and biogeochemical cycle such as carbon and sulfur in the environment [24,65]. Anoxic sulfate-rich environmental conditions such as sea water and ocean salt water promote the survival capacity of SRB. Whereas, in freshwater ambient, the anaerobic metabolic activity would go in minimal level [19]. In the present study, we showed the presence of Sulfate reducing bacteria related to biocorrosion in water and sediment samples from Sisal Port in the Yucatan Peninsula of Mexico.
Our study showed that the optimum temperature was available in most of the sampling sites and other physicochemical parameters also were optimum for the growth of the sulfate reducing bacterial population. Among all theeight genera of SRB present in the samples, Desulfatibacillum and Desulfovibrio were highly abundant in both water and sediment. The reason behind is that out of 37 bacterial genera in SRB group, Desulfovibrio genus is one of the most abundant, opportunists and the second largest in number of species containing genus illustrated as gram-negative bacteria [32,66-68]. In our samples we have found SRB genera such as Desulfobulbus, Desulfotomaculum and Desulfovibrio and their species in the sample next to the gas station according to 16S rRNA gene sequencing analysis; similarly, Desulfovibrio, Desulfotignum and Thermodesulforhabdus sulfate reducing microorganisms (SRM) including archaeal communities as Archaeoglobus genus were found in a high temperature petroleum reservoir by the combined approach of 16S rDNA and 16S rRNA high-throughput sequencing analysis [69]. All eight genera were found in water and sediment of almost all the sampling sites (both freshwater and salt water). Physicochemical parameters did not affect the presence of the bio-corrosion causing SRB in this study.
Laboratory conditions are not similar as in natural environments (Microbiologically-influenced corrosion) for SRB, though various studies have showed that it is possible to culture them under controlled experimental conditions. However, the limitations of the conventional microscopic and culture-based identification techniques allow only a little percentage (less than 1%) of bacteria of the total diversity to be isolated from nature [49,70-72]. Therefore, in this study, 16S rRNA gene sequencing has been used for the identification of sulfate reducing bacteria at the genus level.
Moreover, molecular biology techniques such as Fluorescent in situ hybridization (FISH), microarray DNA technology and functional gene markers such as Dissimilatory sulfite reductase (dsrAB) and adenosine phosphor sulfate reductase (apsA) sequencing analysis could be used for the identification and classification of SRB from different environments. Dsr, APS and Apr play important roles and act as the key enzymes of dissimilatory pathways such as sulfate reduction and sulfur oxidation [73]. Also, dsrAB, apsA and other key sulfate reduction genes are conserved in twenty-five Sulfate-reducing organisms. Further studies are needed to understand the genes involved in biocorrosion mechanisms to control the bio-corrosion caused by Sulfate-reducing bacteria.
Molecular related technologies should develop in order to reduce and recover the huge loss in oil and pipeline industries. Apart from this, it is important to understand the complete mechanism of biocorrosion caused by these ancient bacteria. The dilucidation of the genome sequences of these potent microbes have played a positive role in both industrial and agro-health sectors. Moreover, the exploration of the abundant microorganisms in natural resources could lead to their conservation and a better understanding of their roles in ecological niches. To the best of our knowledge, this is the first 16S rRNA sequence-based report to identify the presence of Sulfate-reducing bacteria related to biocorrosion from environmental water and sediment in the Yucatan peninsula of Mexico.
Conclusion
As a conclusion, our study provides evidence of the presence of SRB in the Yucatan peninsula of Mexico, including the genera Desulfatibacillum, Desulfatitalea, Desulfobacula, Desulfobulbus, Desulfotignum, Desulfotomaculum, Desulfovibrio and Sulfurospirillum which might be involved in biocorrosion mechanisms in various environments. Further genetical and biochemical characterization of these microorganisms will be necessary to dilucidate their role in biocorrosion specifically in the environments studied.
Conflicts of Interest
The authors have declared that no competing interests exist.
Acknowledgment
The present work was funded by El Laboratorio Nacional de Resiliencia Costera-Consejo Nacional de Ciencia y Tecnología (LANRESC-CONACyT), México, Project number: # 293354 and Proyecto Programa de Apoyo a Proyectos de Investigación e Innovación Tecnológica (PAPIIT), México, Project number: # IT202418. The authors would like to acknowledge and appreciate the Faculty of Sciences, Universidad Nacional Autónoma de México, Sisal, Yucatán, México for their continual support and installation facilities. Dr. Muhilan Mahendhiran is thankful for the Postdoctoral scholarship from Dirección General de Asuntos del Personal Académico-Universidad Nacional Autónoma de México (DGAPA-UNAM), Communicating number: 51/2018. Special thanks are due to Lic. Andrés Hiram Zayola Medina from Universidad Nacional Autónoma de México, Sisal, Yucatán, México for his kind help in statistical analyses.
References
- Mori K, Tsurumaru H, Harayama S (2010) Iron corrosion activity of anaerobic hydrogen-consuming microorganisms isolated from oil facilities. J Biosci Bioeng 110: 426-430.
- Hou B, Li X, Ma X, Du C, Zhang D, et al. (2017) The cost of corrosion in China. Npj Materials Degrad 1: 1-10.
- Li X, Zhang D, Liu Z, Li Z, Du C, et al. (2015) Materials science: Share corrosion data. Nature 527: 441-442.
- Guan J, Zhang BL, Mbadinga SM, Liu JF, Gu JD, et al. (2014) Functional genes (dsr) approach reveals similar sulphidogenic prokaryotes diversity but different structure in saline waters from corroding high temperature petroleum reservoirs. Appl Microbiol Biotechnol 98: 1871-1882.
- Mandke JS (1990) Corrosion causes most pipeline failures in Gulf of Mexico. Oil Gas J 88: 40-44.
- Karn SK, Fang G, Duan J (2017) Bacillus sp. acting as dual role for corrosion induction and corrosion inhibition with carbon steel (CS). Front Microbiol 8: 1-11.
- Kruger J (2011) Cost of metallic corrosion: Uhlig’s corrosion handbook. (3rd edtn), Wiley, Hoboken, NJ, USA.
- Little BJ, Lee JS (2014) Microbiologically influenced corrosion: An update. Int Mater Rev 59: 384-393.
- AlAbbas FM, Bhola R, Spear JR, Olson DL, Mishra B (2013) Electrochemical characterization of microbiologically influenced corrosion on linepipe steel exposed to facultative anaerobic Desulfovibrio sp. Int J Electrochem Sci 8: 859-871.
- Javaherdashti R (2008) Microbiologically Influenced Corrosion, an engineering insight. Springer-Verlag London Limited, London, UK.
- Liu H, Gu T, Zhang G, Wang W, Dong S, et al. (2016) Corrosion inhibition of carbon steel in CO2-containing oilfield produced water in the presence of iron-oxidizing bacteria and inhibitors. Corrosion Sci 105: 149-160.
- Dinh HT, Kuever J, Mubmann M, Hassel AW, Stratmann M, et al. (2004) Iron corrosion by novel anaerobic microorganisms. Nature 427: 829-832
- Brown SD, Wall JD, Kucken AM, Gilmour CC, Podar M, et al. (2011) Genome sequence of the mercury-methylation and pleomorphic Desulfovibrio africanus strain Walvis Bay. J Bacteriol 193: 4037-4038.
- Enning D, Garrelfs J (2014) Corrosion of Iron by sulfate-reducing bacteria: New views of an old problem. J Appl Environ Microbiol 80: 1226-1236.
- Beech IB, Gaylarde CC (1999) Recent advances in the study of biocorrosion: An overview. Revista de Microbiologia 30: 177-190.
- Kushkevych I V (2016) Dissimilatory sulfate reduction in the intestinal sulfate-reducing bacteria. Studia Biologica 10: 197-228.
- van den Brand TPH, Roest K, Chen GH, Brdjanovic D, Loosdrecht MCM (2015) Potential for beneficial application of sulfate reducing bacteria in sulfate containing domestic wastewater treatment. World J Microbiol Biotechnol 31: 1675-1681.
- Lens PNL, Kuenen JG (2001) The biological sulfur cycle: Novel opportunities for environmental biotechnology. Water Air Soil Pollut 44: 57-66.
- Colin Y, Goni-Urriza M, Gassie C, Carlier E, Monperrus M, et al. (2017) Distribution of sulfate-reducing communities from estuarine to marine bay waters. Microbio Aquat Sys 73: 39-49.
- Pester M, Knorr KH, Friedrich MW, Wagner M, Loy A (2012) Sulfate-reducing microorganisms in wetlands-fameless actors in carbon cycling and climate change. Front Microbiol 3: 1-19.
- Castro HF, Williams NH, Ogram A (2000) Phylogeny of sulfate-reducing bacteria. FEMS Microbiol Ecol 31: 1-9.
- Kharrat H, Karray F, Bartoli M, Hnia BW (2017) Desulfobulbus aggregans sp. nov., a novel sulfate reducing bacterium isolated from marine sediment from the Gulf of Gabes. Curr Microbiol 74: 449-454.
- Luptakova A (2007) Importance of sulphate-reducing bacteria in environmental biofilms. Nova Biotechnologica VII (I): 17-22.
- Jorgensen BB (1982) Mineralization of organic matter in the seabed, the role of sulphate reduction. Nature 296: 643-645.
- Sánchez-Andrea I, Stams AJM, Amils R, Sanz L (2013) Enrichment and isolation of acidophilic sulfate-reducing bacteria from Tinto River sediments. Environ Microbiol Rep 5: 672-678.
- Jorgensen B, Postgate J (1982) Ecology of the bacteria of the sulphur cycle with special reference to anoxicoxic interface environments. Philos Trans Royal Soc B 298: 543-561.
- Kovac J, Vitezova M, Kushkevych I (2018) Metabolic activity of sulfate-reducing bacteria from rodents with colitis. Open Med 13: 344-349.
- Carbonero F, Benefiel AC, Alizadeh-Ghamsari AH, Gaskins HR (2012) Microbial pathways in colonic sulfur metabolism and links with health and disease. Front Physiol 3: 1-11.
- Marietou A, Roy H, Jorgensen BB, Kjeldsen KU (2018) Sulfate transporters in dissimilatory sulfate reducing microorganisms: A Comparative genomics analysis. Front Microbiol 9: 1-21.
- Muller AL, Kjeldsen KU, Rattei T, Pester M, Loy A (2015) Phylogenetic and environmental diversity of DsrAB-type dissimilatory (bi) sulfite reductases. The ISME J 9: 1152-1165.
- Bradley AS, Leavitt WD, Johnston DT (2011) Revisiting the dissimilatory sulfate reduction pathway. Geobiol 9: 446-457.
- Rabus R, Venceslau SS, Wohlbrand L, Voordouw G, Wall JD, et al. (2015) A post-genomic view of the ecophysiology, catabolism and biotechnological relevance of sulphate-reducing prokaryotes. Adv Microb Physiol 66: 55-321.
- Koschorreck M (2008) Microbial sulphate reduction at a low pH. FEMS Microbiol Ecol 64: 329-342.
- Vallero MVG (2003) Sulfate reducing processes at extreme salinity and temperature: extending its application window. Doctoral Thesis, Wageningen University, Wageningen, the Netherlands.
- Williamson AJ, Carlson HK, Kuehl JV, Huang LL, Lavarone AT, et al. (2018) Dissimilatory sulfate reduction under high pressure by Desulfovibrio alaskensis G20. Front Microbiol 9: 1-10.
- Suzuki D, Ueki A, Amaishi A, Ueki K (2008) Desulfoluna butyratoxydans gen. nov., sp. nov., a novel Gram-negative, butyrate-oxidizing, sulfate-reducing bacterium isolated from an estuarine sediment in Japan. Int J Syst Evol Microbiol 58: 826-832.
- Sherry A, Gray ND, Ditchfield AK, Aktken CM, Jones DM, et al. (2013) Anaerobic biodegradation of crude oil under sulphate-reducing conditions leads to only modest enrichment of recognized sulphate-reducing taxa. Int Biodeterior Biodegradation 81: 105-113.
- Wang R (2012) Physiological implication of hydrogen sulfide: A whiff exploration that blossomed. Physiol Rev 92: 791-896.
- Karlin S, Brocchieri L, Mrázek J, Kaiser D (2006) Distinguishing features of delta-proteobacterial genomes. Proc Natl Acad Sci U S A 103: 11352-11357.
- Mori K, Kim H, Kakegawa T, Hanada S (2003) A novel lineage of sulfate-reducing microorganisms: Thermodesulfobiaceae fam. nov., Thermodesulfobium narugense, gen. nov., sp. nov., a new thermophilic isolate from a hot spring. Extremophiles 7: 283-290.
- Rojas-Herrera R, Narváez-Sapata J, Zamudio-Maya M, Mena-Martínez ME (2008) A simple silica-based method for metagenomic DNA extraction from soil and sediments. Mol Biotechnol 40: 13-17.
- Zhang J, Kobert K, Flouri T, Stamatakis A (2014) PEAR: A fast and accurate Illumina paired-end read merger. Bioinformatics 30: 614-260.
- Schmieder R, Edwards R (2011) Quality control and preprocessing of metagenomic datasets. BMC Bioinformatics 27: 863-864.
- Schmieder R, Lim YW, Rohwer M, Edwards R (2010) TagCleaner: Identification and removal of tag sequences from genomic and metagenomic datasets. BMC Bioinformatics 11: 341.
- Minot SS, Krumm N, Greenfield NB (2015) One Codex: A sensitive and accurate data platform for genomic microbial identification. bioRxiv. 027607. 1-23.
- Lever J, Krzywinski M, Altman N (2017) Points of significance: Principal component analysis. Nature Methods 14: 641-642.
- Zhang Z, Murtagh F, Poucke SV, Lin S, Lan P (2017) Hierarchical cluster analysis in clinical research with heterogeneous study population, highlighting its visualization with R. Ann Transl Med 5: 1-11.
- Li Y, Xu D, Chen C, Li X, Jia R, et al. (2018) Anaerobic microbiologically influenced corrosion mechanisms interpreted using bioenergetics and bio electrochemistry: A review. J Mater Sci Technol 34: 1713-1718.
- Gu T, Xu D, Zhang P, Li Y, Lindenberger AL (2015) Microbiologically Influenced Corrosion and Its Impact on Metals and Others Materials: Microbiology for Minerals, Metals, Materials and Environment. CRC Press, Boca Raton, Florida, USA.
- Promnuan K, O-Thong S (2017) Biological hydrogen sulfide and sulfate removal from rubber smoked sheet wastewater for enhanced biogas production. Energy Procedia 138: 569-574.
- Muyzer G, Stams AJM (2008) The ecology and biotechnology of sulphate-reducing bacteria. Nat Rev Microbiol 6: 441-454.
- Thauer RK, Stackebrandt E, Hamilton WA (2007) Energy metabolism and phylogenetic diversity of sulphate-reducing bacteria. Sulphate-Reducing Bacteria: Environmental and Engineered Systems. Cambridge University Press, Cambridge, UK.
- Sorokin DY, Tourova TP, Henstra AM, Stams AJM, Galinski EA, et al. (2008) Sulfidogenesis under extremely haloalkaline conditions by Desulfonatronospira thiodismutans gen. nov., sp. nov., and Desulfonatronospira delicata sp. nov.: A novel lineage of Deltaproteobacteria from hypersaline soda lakes. Microbiol 154: 1444-1453.
- Zhilina T, Zavarzin G, Rainey F, Pikuta E, Osipov G, et al. (1997) Desulfonatronovibrio hydrogenovorans gen. nov., sp. nov., an alkaliphilic, sulfate-reducing bacterium. Int J Syst Evol Microbiol 47: 144-149.
- Barton LL, Fauque GD (2009) Biochemistry, physiology and biotechnology of sulfate-reducing bacteria. Adv Appl Microbiol 68: 41-98.
- Visser M, Stams AJM, Frutschi M, Bernier-Latmani R (2016) Phylogenetic comparison of Desulfotomaculum species of subgroup 1a and description of Desulfotomaculum reducens sp. nov. nt. J Syst Evol Microbiol 66: 762-767.
- Aullo T, Ranchou-Peyruse A, Ollivier B, Magot M (2013) Desulfotomaculum spp. and related gram-positive sulfate-reducing bacteria in deep subsurface environments. Front Microbiol 4: 1-12.
- Visser M, Worm P, Muyzer G, Pereira IAC (2013) Genome analysis of Desulfotomaculum kuznetsovii strain 17T reveals a physiological similiarity with Pelotomaculum thermopropionicum strain SIT. Stand Genomic Sci 8: 69-87.
- Visser M, Parshina SN, Alves JI, Sousa DZ, Pereira IAC, et al. (2014) Genome analyses of the carboxydotrophic sulfate-reducers Desulfotomaculum nigrificans and Desulfotomaculum carboxydivorans and reclassification of Desulfotomaculum caboxydivorans as a later synonym of Desulfotomaculum nigrificans. Stand Genomic Sci 9: 655-675.
- Ehrlich HL, Newman DK (2009) Geomicrobiology. (5th edtn), CRC Press, New York, NY, USA.
- Badziong W, Thauer RK (1978) Isolation and characterization of desulfovibrio growing on hydrogen plus sulfate as the sole energy source. Arch Clin Microbiol 116: 41-49.
- Feio MJ, Zinkevich V, Beech IB, Llobet-Brossa E, Eaton P, et al. (2004) Desulfovibrio alaskensis sp. nov., a sulphate-reducing bacterium from a sourced oil reservoir. Int J Syst Evol Microbiol 54: 1747-1752.
- Plugge CM, Zhang W, Scholten JCM, Stams AJM (2011) Metabolic flexibility of sulfate-reducing bacteria. Front Microbiol 2: 1-8.
- Robador A, Muller AL, Sawicka JE, Berry D, Hubert CRJ, et al. (2016) Activity and community structures of sulfate-reducing microorganisms in polar, temperate and tropical marine sediments. The ISME J 10: 796-809.
- Crispim JS, Dias RS, Vidigal PMP, Paula de Sousa M, Canedo da Silva C, et al. (2018) Screening and characterization of prophages in Desulfovibrio genomes. Sci Rep 8: 1-10.
- Dar SA, Kleerebezem R, Stams AJM, Kuenen G, Muyzer G (2008) Competition and coexistence of sulfate-reducing bacteria, acetogens and methanogens in a lab-scale anaerobic bioreactor as affected by chaning substrate to sulfate ratio. Appl Microbiol Biotechnol 78: 1045-1055.
- Purdy KJ, Nedwell DB, Embley TM, Takii S (1997) Use of 16S rRNA-targeted oligonucleotide probes to investigate the occurrence and selection of sulfate-reducing bacteria in response to nutrient addition to sediment slurry microcosms from a Japanese estuary. FEMS Microbiol Ecol 24: 221-234.
- Li XX, Liu JF, Zhou L, Mbadinga SM, Yang SZ, et al. (2017) Diversity and composition of sulfate-reducing microbial communities based on genomic DNA and RNA transcription in production water of high temperature and corrosive oil reservoir. Front Microbiol 8: 1-17.
- Pinevich AV, Andronov EE, Pershina EV, Pinevich AA, Dmitrieva HY (2018) Testing culture purity in prokaryotes: Criteria and challenges. Antonie van Leeuwenhoek 111: 1509-1521.
- Stewart EJ (2012) Growing Unculturable Bacteria. J Bacteriol 194:16. 4151-4160.
- Rooney-Varga JN, Devereux R, Evans RS, Hines ME (1997) Seasonal changes in the relative abundance of uncultivated sulfate-reducing bacteria in a salt marsh sediment and in the rhizosphere of Spartina alterniflora. J Appl Environ Microbiol 63: 3895-3901.
- Meyer B, Kuever J (2008) Homology modeling of dissimilatory APS reductases (AprBA) of sulfur-oxidizing and sulfate reducing prokaryotes. PLoS One 3: e1514.
- Pereira IAC, Ramos AR, Greing F, Marques MC, de Silva SM, et al. (2011) A comparative genomic analysis of energy metabolism in sulfate reducing bacteria and archaea. Front Microbiol 2: 1-22.
 Spanish
Spanish  Chinese
Chinese  Russian
Russian  German
German  French
French  Japanese
Japanese  Portuguese
Portuguese  Hindi
Hindi