Review Article, J Plant Physiol Pathol Vol: 6 Issue: 4
DNA Methylation and Plant Stress Responses
Jianchuan Deng1#, Shuyan Kou2#, Qian Zou2, Ping Li3,4*, Cuiping Zhang5 and Pingrong Yuan2*
1Institute of Economical Crops, Yunnan Academy of Agricultural Sciences, Kunming, Yunnan, China
2Institute of Foodl Crops, Yunnan Academy of Agricultural Sciences, Kunming, Yunnan, China
3University of Chinese Academy of Sciences, Beijing, China
4Key Laboratory of Economic Plants and Biotechnology, Kunming Institute of Botany, Chinese Academy of Sciences, Yunnan Key Laboratory for Wild Plant Resources, Kunming, Yunnan, China
5YuXi Academy of Agricultural Sciences, Yuxi, China
#These authors contributed equally
*Corresponding Author : Ping Li
Key Laboratory of Economic Plants and Biotechnology, Kunming Institute of Botany, Chinese Academy of Sciences
Yunnan Key Laboratory for Wild Plant Resources, Kunming, Yunnan 650201, China
E-mail: yuanpr2003@aliyun.com
Pingrong Yuan
Institute of Food Crops, Yunnan Academy of Agricultural Sciences, Kunming, Yunnan, 650205, China
E-mail: liping@mail.kib.ac.cn
Received: December 29, 2016 Accepted: March 09, 2017 Published: March 13, 2017
Citation: Deng J, Kou S, Zhang C, Zou Q, Li P, et al. (2018) DNA Methylation and Plant Stress Responses. J Plant Physiol Pathol 6:4. doi: 10.4172/2329-955X.1000182
Abstract
Plants are sensitive to external environmental conditions and the mechanisms of stress tolerance in plants are complex. As one of the most significant epigenetic phenomena, DNA methylation plays a crucial role in plants’ adaptation to environmental changes. Recent research has improved our understanding of the relationship between DNA methylation and stress tolerance. Here, we review the mechanism of DNA methylation and discuss recent reports of the dynamics of DNA methylation in plants under abiotic stress conditions, including high salt concentration, drought, and extreme temperatures. The DNA methylation plays important functions during the response to stress in plants, but the differential variations of which were exhibiting in different species. The consistent or the specific mechanisms of DNA methylation need further exploration.
Keywords: DNA Methylation; Abiotic Stress; RNA-Directed DNA Methylation (Rddm); Small RNA
Introduction
Epigenetic mechanisms change gene expression without altering DNA sequences, and mainly include DNA methylation, genomic imprinting, gene silencing, chromatin modifications and RNA editing. As an epigenetic marker, DNA methylation has received increasing attention in recent years. All nucleotides in a given DNA sequence can be methylated, but the most pervasive and important form is cytosine methylation [1]. In plants, DNA methylation, usually found in repetitive regions, is divided into three types according to the base following the methylated cytosine; CG, CHG, or CHH (where H represents A, C or T) [2]. DNA methylation can be achieved in two ways: de novo methylation and maintenance [3].
As plants are sessile, they are more susceptible to damage than animals under biotic and abiotic stress. abiotic stresses, including high salinity, high or low temperatures, and drought, cause serious losses in agricultural productivity each year. In recent years, a growing number of studies have indicated that methylation status changes frequently (especially in stress-related genes) in developing plants under abiotic stress [4,5]. This review briefly summarizes the mechanism by which DNA is methylated and describes changes in DNA methylation observed in plants affected by abiotic stress.
Mechanism of RNA-directed DNA methylation
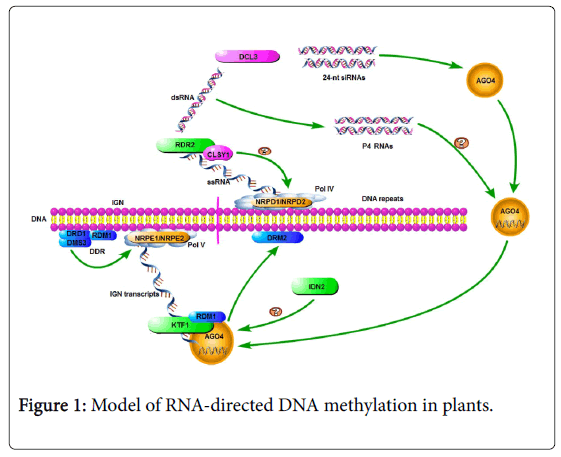
RNA-directed DNA methylation (RdDM) is a pathway first discovered in plants by which DNA is methylated. Most elements of the RdDM pathway have been elucidated (Figure 1). DNA-dependent plant-specific RNA polymerase IV (Pol IV) and Pol V are vital in RdDM; their largest subunits are NRPD1 and NRPE1, respectively [6,7]. Pol IV uses RdDM target loci as templates to synthesize abnormal single-stranded RNA (ssRNA) transcripts, and Pol V transcribes intergenic non-coding (IGN) regions into IGN transcripts in the following process.
CLASSY1 (CLSY 1) is a chromatin remodeling factor that recruits Pol IV and facilitates its transcription activity [8]. RNA-DEPENDENT RNA POLYMERASE 2 (RDR2) generates double-stranded RNA (dsRNA) from ssRNA [9]. DICER-LIKE 3 proteins (DCL3) cleaves dsRNA into 24-nucleotide (nt) small interfering RNAs (siRNAs) [9]. These 24-nt siRNAs are bound by the ARGONAUTE (AGO) proteins, then colocalize with IGN transcripts produced by Pol V [10,11]. The putative chromatin remodeling factors DEFECTIVE IN RNADIRECTED DNA METHYLATION 1 (DRD1), DEFECTIVE IN MERISTEM SILENCING 3 (DMS3) and RNA-DIRECTED DNA METHYLATION 1 (RDM1) form a stable protein complex – DDR – in the RdDM pathway. This complex is indispensable in Pol V transcription, and combines with chromatin. IGN transcripts can recruit AGO4 [12]. INVOLVED IN DE NOVO 2 (IDN2) is a SUPPRESSOR OF GENE SILENCING 3 (SGS3)-like protein, and stabilizes the interactions between siRNAs and IGN transcripts [12]. In plants, IGN transcripts, AGO4, RDM1 and KTF1 form a complex that directs the DNA methyltransferase DRM2 to target loci [12].
Previous studies shown that small RNAs, especially 24-nt siRNAs, are pivotal in the RdDM pathway [13-16]. But not all methylated target loci depend on dicer activity that produces 24-nt siRNAs [17,18]. The discovery of a new class of Pol IV-dependent RNAs (‘P4 RNAs’) suggests that most RdDM loci maintain methylation levels through P4 RNAs; these loci are not or partially dependent on Dicer-like protein or 24-nt siRNAs (Figure 1) [18,19]. During RdDM pathway, P4 RNAs transformed to double stranded RNA with the function of RDR2 (RNA- dependent RNA polymerase 2), from which 24-nt siRNAs can got by DCL3 (Dicer-like 3) [20,21]. The then single-stranded RNA the P4 RNAs in the size range of ~25-50 nt are produced by Pol IV and precursors of 24-nt siRNAs [18].
Abnormal single-stranded RNAs (ssRNA) are generated by RNA Polymerase IV (Pol IV) in DNA repeat regions. CLSY1 recruits Pol IV and facilitates its transcription. RDR2 converts the abnormal ssRNAs to double-stranded RNAs (dsRNAs), which are then trimmed into 24- nt small interfering RNAs (siRNAs) by DCL3. The 24-nt siRNAs are bound by AGO4, which then interacts with NRPE1. When 24-nt siRNAs are bound to AGO4, KTF1 binds the IGN transcripts produced by Pol V, and combines with AGO4 by base-pairing between siRNA and IGN transcripts. DRD1, DMS3 and RDM1 form a complex to stabilize the interaction between NRPE1 and chromatin. IDN2 recognizes this complex and stabilizes the interaction between siRNAs and IGN transcripts.
Methylation level changes under salt stress
Salinity is one of the most serious environmental limiting factors for crop productivity [22]. Adverse effects of high salt concentration include decreased plant cell activity, and decreased productivity, germination rate and plant height [23,24]. The methylation status of different species and varieties varies under salt stress [25,26]. For example, the DNA methylation levels of rice variety ‘IR29 ’ and rapeseed variety ‘Exagone’ both decrease under salt stress. Interestingly, when Exagone plants were grown under conditions of salt stress for two weeks, the number of fully methylated bands increased in the plants allowed to recover from salt stress for 1-2 days compared with plants grown in high salinity for the entire period [23,27] During the salt stress, there are clear changes in the methylation of genes involved in physiological processes in Exagone; for example, the polymorphic fragment Bn_05 is homologous to the gene encoding Plasma Membrane Intrinsic Protein 2, which is involved in salt tolerance, and the polymorphic fragment Bn_09 is homologous to an Arabidopsis thaliana gene that encodes a Trehalose Phosphatase/ Synthase 4 (TPS4) protein, which is involved in trehalose biosynthesis. Trehalose, which is a nonreducing disaccharide of glucose, can increase plants stress tolerance through production of carbohydrate alterations [28].
Methylation status can also differ within a species. In cotton, the methylation level of salt-tolerant cultivars ‘CCRI 35’ and ‘Zhong 07’ decreases during salt stress, while the methylation ratio of salt-sensitive ‘CCRI 12’ increases from 68.1% to 75.8% [29]. One DNA fragment with an altered methylation level is found to be homologous to a gene encoding a MYB transcription factor that is involved in stress tolerance [30]. In Arabidopsis thaliana, a transcription factor, the R2R3-MYB gene, was transcriptionally regulated through RdDM under salt stress. It is found that a region about 500 bp upstream region of AtMYB74 has a heavily reduced methylated level in respons to high sanity [31]. In rice, methylation decreased in the root and shoot of ‘Geumgangbyeo’ and ‘IR29’ under salinity stress, whereas it increased in ‘Nip’ and ‘POK’ [32]. The number of methylated bands decreased in root tissue of all genotypes, especially Nip [33].
Methylation level changes under drought stress
Drought is the most serious environment stress that limits plant development and production in most rain-fed systems worldwide [34,35]. In recent decades, drought stress-related plant mortality has increased [36]. Plant height, flowering, ratio of spikelet sterility, and leaf shape and color are also affected by drought stress [37]. Plants under drought stress exhibit dynamic methylation levels [38-40]. In Populus trichocarpa, drought stress increases methylation; using bisulfite sequencing (BS-seq) to detect methylation levels, the proportion of methylated cytosines is 10.04% under drought stress, while it is only 7.75% in the well-watered treatment [41]. A positive correlation was found between gene expression and methylation levels under drought stress, demonstrated by 7329 upregulated genes with increased methylation levels, and 10 322 downregulated genes with decreased methylation levels [41]. The same pattern was noted in the DNA methylation levels of the drought-susceptible rice varieties IR20 and CO43: methylation levels increase under drought [42]. In barley, the genome-wide DNA methylation level is high and stable under water-deficiency conditions, but the methylation sites showed difference between leaf and roots [43]. The repetitive elements show a de-methylated tendency in leaves while methylated in roots. Moreover, majority of stress-induced methylation events would be reversible and the barley plant would recover [43].
There are also different changes in methylation status in other species or genotypes under drought stress. For example, using MSAP (Methylation-Sensitive Amplification Polymorphism), the methylation level of perennial ryegrass reduced from 57.67% in the control to 47.39% under drought stress; the level of fully methylated DNA is notably down-regulated by 12.58% in the drought condition [44]. One fragment in which the methylation level decreases under drought stress is found to be homologous to LOC100833067, which encodes respiratory burst oxidase homolog (Rboh) protein [44]. Rboh-NADPH oxidases produce oxygen intermediates that regulate abiotic stress responses [45]. In the drought-tolerant rice varieties ‘PMK3’ and ‘PL’, methylation levels also decreased under drought stress compared with the control condition [42]. Interestingly, the methylation patterns in rice (Oryza sativa L.) are affected by multi-generational drought stress treatment [37,42]. In order to adapt to a long term drought stress, rice perform a high proportion of drought-induced epimutations through altered DNA methylation status [46].
Methylation level changes under temperature extremes
In the frigid Zone and at high altitudes, and in the tropics, extreme temperatures affect plant growth, including cell growth, plant height, coleoptile growth, shoot dry mass, crop yield and flavor [47-49]. As under other abiotic stresses, changes in methylation level are dynamic under temperature stress [50]. Compared with normal temperature, the methylation rate of cotton (Gossypium hirsutum) decreases under cold stress, and continues to decrease with prolonged cold treatment [29]. The methylation rate of cotton seedlings recovers somewhat when subsequently grown under a normal temperature, but never returns to its original level [51]. Two fragments shown to be demethylated under cold stress were found to be homologous to G. hirsutum transcription factors. One, WRKY1, belongs to a gene family involved in plant defense [52], while the other, MYB, belongs to a superfamily of transcription regulatory proteins with key roles in plant development and responses to abiotic stress [53]. The methylation rate of maize (Zea mays L.) also decreases under cold stress [54]. In this species, the polymorphic fragment DM 113 is found to be homologous to LOC100281677, which encodes cystathionine β -synthase (CBS) domain-containing protein. The CBS domain has no defined functions but is involved in the catalytic activity of many enzymes [55]. In tobacco, overexpression of CBS domain-containing protein can improve salinity, oxidative, and heavy metal tolerance [56].
In Populus simonii, methylation levels decrease under heat stress, and increase under cold conditions. To investigate DNA methylation changes under abnormal temperatures, three groups of 10 plants were exposed to 42°C, 4°C and 25°C for 3, 6, 12 and 24 h, respectively. Using MSAP to assess DNA methylation levels, the lowest levels of DNA methylation in P. simonii were found in the high temperature condition, and the highest levels under the coldest temperature condition [57]. The short-term heat shock will trigger the DNA methylation in cultured microspores of Brassica napus cv . Topas, through which the methylated region-related genes (DRGs) were about twice higher than that of 18°C for 6 h, after treatment at 32°C for 6 h [58].
There is an interesting phenomenon that DNA methylation changes contributed to cross-adaptation process. In Brassica rapa , the cold induced methylation would confer a heat tolerance. 1562 differentially methylated genes are indentified in cold treatment Cold acclimation alters DNA methylation patterns and confers tolerance to heat and increases growth rate in Brassica rapa., during which four methylated genes, including BramMDH1, BraKAT2, BraSHM4, and Bra4CL2, are related to increase the organic acids and photosynthesis levels, and then lead to a improved heat-tolerance and high growth rate [59].
Table 1 shows the polymorphic fragments identified from different species and tissues under control and abiotic stress conditions, and their respective homologous genes, revealed using BLAST. Gene expression is interrelated with gene methylation status, i.e., changes in methylation levels affect gene expression. Some genes, such as Os09g27580, GU324346.1, NM_001157237 and NM_001154597, match with polymorphic fragments that are known to be involved with abiotic response in plants [55,60,61] while others are involved in growth, flowering, and ROS generation [62,63].
| Stress | Species or tissue | Homologous gene | Methylation change (compared with control) | Expression change (compared with control) | Function | Reference |
|---|---|---|---|---|---|---|
| Drought | Perennial ryegrass | LOC100839781 | Demethylated | Up-regulated | B. distachyon peroxidase 12-like, mRNA | [44] |
| LOC100833067 | Demethylated | Up-regulated | B. distachyon respiratory burst oxidase homolog protein B-like, mRNA | |||
| Rice | Os09g27580 | Methylated | Down-regulated | HAK 5, high affinity K+ transporter 5 | [38] | |
| Os10g39130 | Methylated | Down-regulated | AGL19, AGAMOUS-like 19 | |||
| Barley | MLOC_53923 | Methylated | Down-regulated | HvCKX2.1 CYTOKININ-OXIDASE 2.1 |
[40] | |
| Salt | Rice | Os03g44130 | Demethylated | Up-regulated | OsMYB91 R2R3-type MYB gene |
[64] |
| Cotton (CCRI 35) | GU324346.1 | Demethylated | Up-regulated | Theobroma cacao cultivar Scavina 6 TT2 like MYB transcription factor (MYBPA) mRNA, complete cds | [29] | |
| Rice (IR29 shoot) | Os02g52290 | Methylated | Up-regulated | Peptidyl-prolyl cis-trans isomerase, FKBP-type | [23] | |
| Rice (IR29 shoot) | Os02g44330 | Methylated | Down-regulated | Rho guanine nucleotide exchange factor | ||
| Rice (IR29 shoot) | Os03g32490 | Demethylated | Up-regulated | DUF1230 domain-containing | ||
| Rice (IR29 shoot) | Os11g23900 | Demethylated | Down-regulated | Retrotransposon protein | ||
| Temperature | Maize (Zea mays L) | NM_001153998 | Demethylated | Up-regulated | Receptor kinase | [54] |
| NM_001157237 | Demethylated | Up-regulated | Calcium-activated outward-rectifying potassium channel 1 | |||
| NM_001154597 | Demethylated | Up-regulated | CBS domain-containing protein |
Table 1: Gene expressions changed under stress condition.
The methylation status of different species and genotypes varies under abiotic stress, and methylation ratio change can affect gene expression, especially stress-tolerance related genes. Maybe these stress-tolerance related genes perform their function through changing methylation level.
Conclusion
This review briefly introduced the RdDM pathway, which increases DNA methylation. Recent research has shown that not all targeted loci are fully dependent on 24-nt siRNA, and that 25-50 nt P4 RNAs may have a similar function. Changes in methylation levels under abiotic stress reveal that different species and genotypes vary in methylation levels in different ways, and that in some cases there is even variation within species under certain stress conditions.
Despite the fact that recent studies have increased our knowledge of DNA methylation and its relationship to stress responses, the answers to several questions remain unknown; for example, what are the exact functional mechanisms of P4 RNAs? What’s the general mechanism of plant DNA methylation in regulation with gene expression and maintain? Obtaining further insights into methylation and the precise pathway in which it alters DNA expression will be a pivotal challenge for future studies.
Acknowledgements
We thank for Dr. Gaojing Liu correcting the English text, Professor Li Liu and Professor Yasser Zain ElAbden Elrefaee for his suggestion on this paper. This work was supported by Personnel Training Program of Academic and Technical Leaders of Yunnan Province, Yunnan Rice Product System (2018KJTX001), and Main Project for Biological Seed Industry of Yunnan Province (2015ZA003).
References
- Rabinowicz PD, Schutz K, Dedhia N, Yordan C, Parnell LD, et al. (1999) Differential methylation of genes and retrotransposons facilitates shotgun sequencing of the maize genome. Nat Genet 23: 305-308.
- Tan F, Zhou C, Zhou Q, Zhou S, Yang W, et al. (2016) The ortholog of ddm1 is mainly required for chg and cg methylation of heterochromatin and is involved in drm2-mediated chh methylation that targets mostly genic regions of the rice genome. Plant Physiol 171: 1-44.
- Hitt MM, Wu TL, Cohen G, Linn S (1988) De novo and maintenance dna methylation by a mouse plasmacytoma cell dna methyltransferase. J Biol Chem 263: 4392.
- Brzezinka K, Altmann S, Czesnick H, Nicolas P, Gorka M, et al. (2016) Arabidopsis FORGETTER1 mediates stress-induced chromatin memory through nucleosome remodeling. Elife 5: e17061.
- Colicchio JM, Miura F, Kelly JK, Ito T, Hileman LC (2015) DNA methylation and gene expression in Mimulus guttatus. BMC Genom 16: 507.
- Huettel B, Kanno T, Daxinger L, Bucher E, van der Winden J, et al. (2007) RNA-directed DNA methylation mediated by DRD1 and Pol IVb: A versatile pathway for transcriptional gene silencing in plants. Biochim Biophys Acta 1769: 358-374.
- Pikaard CS, Haag JR, Ream T, Wierzbicki AT (2008) Roles of RNA polymerase IV in gene silencing. Trends Plant Sci 13: 390-397.
- Smith L, Pontes O, Searle L, Yelina N, Yousafzai F, et al. (2007) An snf2 protein associated with nuclear rna silencing and the spread of a silencing signal between cells in arabidopsis. Plant Cell 19: 1507-1521.
- Xie Z, Johansen LK, Gustafson AM, Kasschau KD, Lellis AD, et al. (2004) Genetic and functional diversification of small RNA pathways in plants. PLoS Biol 2: E104.
- Li CF, Pontes O, El-Shami M, Henderson IR, Bernatavichute YV, et al. (2006) An ARGONAUTE4-containing nuclear processing center colocalized with Cajal bodies in Arabidopsis thaliana. Cell 126: 93-106
- Pontes O, Li CF, Costa Nunes P, Haag J, Ream T, et al. (2006) The Arabidopsis chromatin-modifying nuclear siRNA pathway involves a nucleolar RNA processing center. Cell 126: 79-92.
- Ausin I, Mockler TC, Chory J, Jacobsen SE (2009) IDN1 and IDN2 are required for de novo DNA methylation in Arabidopsis thaliana. Nat Struct Mol Biol 16: 1325-1327.
- Lewsey MG, Hardcastle TJ, Melnyk CW, Molnar A, Valli A, et al. (2016) Mobile small RNAs regulate genome-wide DNA methylation. Proc Natl Acad Sci USA 113: 801-810.
- Xu W, Yang T, Dong X, Li DZ, Liu A (2016) Genomic DNA methylation analyses reveal the distinct profiles in Castor bean seeds with persistent endosperms. Plant Physiol 171: 1242-1258.
- Grover JW, Kendall T, Baten A, Burgess D, Freeling M, et al. (2018) Maternal components of RNAâ€Âdirected DNA methylation are required for seed development in Brassica rapa. Plant J 94: 575-582.
- LiuW, Duttke SH, Hetzel J, Groth M, Feng S, et al. (2018) RNA-directed DNA methylation involves co-transcriptional small-RNA-guided slicing of polymerase V transcripts in Arabidopsis. Nat Plants 4: 181-188.
- Fei Y, Xue Y, Du P, Yang S, Deng X (2016) Expression analysis and promoter methylation under osmotic and salinity stress of TaGAPC1 in wheat (Triticum aestivum L). Protoplas 254: 987-996.
- Yang DL, Zhang G, Tang K, Li J, Yang L, et al. (2016) Dicer-independent RNA-directed DNA methylation in Arabidopsis. Cell Res 26: 66-82.
- Dalakouras A, Dadami E, Wassenegger M, Krczal G, Wassenegger M (2016) RNA-directed DNA methylation efficiency depends on trigger and target sequence identity. Plant J 87: 202-214.
- Law JA, Jacobsen SE (2010) Establishing, maintaining and modifying dna methylation patterns in plants, and animals. Nat Rev Genet 11: 204-220.
- Matzke MA, Mosher RA (2014) Rna-directed dna methylation: An epigenetic pathway of increasing complexity. Nat Rev Genet 15: 394-408.
- Vadawale A, Mihani R, Mathew A, Robin P (2012) Transformation of groundnut - arachis hypogea l. var. gg20 with the cox gene-an attempt to develop salinity tolerance. Int J Pharma Bio Sci 3: B591-B599.
- Karan R, DeLeon T, Biradar H, Subudhi PK (2012) Salt stress induced variation in DNA methylation pattern and its influence on gene expression in contrasting rice genotypes. PLoS One 7: e40203.
- Wang W, Zhao X, Pan Y, Zhu L, Fu B, et al. (2011) DNA methylation changes detected by methylation-sensitive amplified polymorphism in two contrasting rice genotypes under salt stress. J Genet Genom 38: 419-24.
- Paul A, Dasgupta P, Roy D, Chaudhuri S (2017) Comparative analysis of Histone modifications and DNA methylation at OsBZ8 locus under salinity stress in IR64 and Nonabokra rice varieties. Plant Mol Biol 95: 63-88.
- Yaish MW, Al-Lawati A, Al-Harrasi I, Patankar HV (2018) Genome-wide DNA Methylation analysis in response to salinity in the model plant caliph medic (Medicago truncatula). BMC Genom 19: 78.
- Marconi G, Pace R, Traini A, Raggi L, Lutts S, et al. (2013) Use of MSAP markers to analyse the effects of salt stress on DNA methylation in rapeseed (Brassica napus var. oleifera). PLoSOne 8: e75597.
- Cortina C, Culianezmacia FA (2005) Tomato abiotic stress enhanced tolerance by trehalose biosynthesis. Plant Sci 169: 75-82.
- Wang B, Fu R, Zhang M, Ding Z, Chang L, et al. (2015) Analysis of methylation-sensitive amplified polymorphism in different cotton accessions under salt stress based on capillary electrophoresis. Genes & Genomics 37: 713-724.
- Agarwal M, Hao Y, Kapoor A, Dong CH, Fujii H, et al. (2006) A R2R3 type MYB transcription factor is involved in the cold regulation of CBF genes and in acquired freezing tolerance. J Biol Chem 281: 37636-37645.
- Xu R, Wang Y, Zheng H, Lu W, Wu C, et al. (2015) Salt- induced transcription factor MYB74 is regulated by the RNA- directed DNA methylation pathway in Arabidopsis. J Exp Bot 66: 5997-6008.
- Karan R, Deleon T, Biradar H, Subudhi P (2005) Epigenetic influence on gene expression under salinity stress in salt tolerant and salt susceptible rice cultivars. Int Plant Animal Genome Conf 485: 252-256.
- Zeng FS, Li LL, Liang NS, Wang X, Li X, et al. (2015) Salt tolerance and alterations in cytosine methylation in the interspecific hybrids of Fraxinus velutina and Fraxinus mandshurica. Euphytica 205: 721-737.
- Adams HD, Guardiola-Claramonte M, Barron-Gafford GA, Villegas JC, Breshears DD, et al. (2009). Temperature sensitivity of drought-induced tree mortality portends increased regional die-off under global-change-type drought. Proc Natl Acad Sci USA 106: 7063-7066.
- McDowell N, Pockman WT, Allen CD, Breshears DD, Cobb N, et al. (2008) Mechanisms of plant survival and mortality during drought: Why do some plants survive while others succumb to drought?. New Phytol 178: 719-739.
- Allen CD, Macalady AK, Chenchouni H, Bachelet D, McDowell N, et al. (2010) A global overview of drought and heat-induced tree mortality reveals emerging climate change risks for forests. Forest Eco Manag 259: 660-684.
- Wang WS, Pan YJ, Zhao XQ, Dwivedi D, Zhu LH, et al. (2011) Drought-induced site-specific DNA methylation and its association with drought tolerance in rice (Oryza sativa L.). J Exp Bot 62: 1951-1960.
- Garg R, Narayana Chevala V, Shankar R, Jain M (2015) Divergent DNA methylation patterns associated with gene expression in rice cultivars with contrasting drought and salinity stress response. Sci Rep 5: 14922.
- Ganguly DR, Crisp PA, Eichten SR, Pogson BJ (2017) The Arabidopsis DNA methylome is stable under transgenerational drought stress. Plant Physiol 175: 1893-1912.
- Surdonja K, Eggert K, Hajirezaei MR, Harshavardhan VT, Seiler C, et al. (2017) Increase of DNA methylation at the HvCKX2.1 promoter by terminal drought stress in barley. Epigenomes 1: 9.
- Liang D, Zhang ZJ, Wu HL, Huang CY, Shuai P, et al. (2014) Single-base-resolution methylomes of populus trichocarpa reveal the association between DNA methylation and drought stress. Bmc Genetics 15: 1-9.
- Gayacharan, Joel AJ (2013) Epigenetic responses to drought stress in rice (Oryza sativa L.). Physiol Mol Biol Plants 19: 379-387.
- Chwialkowska K, Nowakowska U, Mroziewicz A, Szarejko I, Kwasniewski M (2016) Water- deficiency conditions differently modulate the methylome of roots and leaves in barley (Hordeum vulgare L.). J Exp Bot 67: 1109-1121.
- Tang XM, Tao X, Wang Y, Ma DW, Li D, et al. (2014) Analysis of DNA methylation of perennial ryegrass under drought using the methylation-sensitive amplification polymorphism (MSAP) technique. Mol Genet Genomics 289: 1075-1084.
- Torres MA, Dangl JL (2005) Functions of the respiratory burst oxidase in biotic interactions, abiotic stress and development. Curr Opin Plant Biol 8: 397-403.
- Zheng XG, Liang C, Hui X, Haibin W, Qiaojun L, et al. (2017) Transgenerational epimutations induced by multi- generation drought imposition mediate rice plant’s adaptation to drought condition. Sci Rep 7: 39843.
- Bañon S, Fernandez JA, Franco JA, Torrecillas A, Alarcón JJ, et al. (2004) Effects of water stress and night temperature preconditioning on water relations and morphological and anatomical changes of Lotus creticus plants. Scien Horticul 101: 333-342.
- Centomani I, Sgobba A, D'Addabbo P, Dipierro N, Paradiso A, et al. (2015) Involvement of DNA methylation in the control of cell growth during heat stress in tobacco BY-2 cells. Protoplasma 252: 1451-1459.
- Zhang B, Tieman DM, Jiao C, Xu Y, Chen K, et al. (2016) Chilling-induced tomato flavor loss is associated with altered volatile synthesis and transient changes in DNA methylation. Proc Natl Acad Sci USA 113: 15580-12585.
- Yu Y, Yang X, Wang H, Shi F, Liu Y, et al. (2013) Cytosine methylation alteration in natural populations of Leymus chinensis induced by multiple abiotic stresses. PLoSOne 8: e55772.
- Fan HH, Wei J, Li TC, Li ZP, Guo N, et al. (2013) DNA methylation alterations of upland cotton (Gossypium hirsutum) in response to cold stress. Acta Physiologiae Plantarum 35: 2445-2453.
- Eulgem T, Somssich IE (2007) Networks of WRKY transcription factors in defense signaling. Curr Opin Plant Biol 10: 366-371.
- Yanhui C, Xiaoyuan Y, Kun H, Meihua L, Jigang L, et al. (2006) The MYB transcription factor superfamily of Arabidopsis: Expression analysis and phylogenetic comparison with the rice MYB family. Plant Mol Biol 60: 107-124.
- Shan X, Wang X, Yang G, Wu Y, Su S, et al. (2013). Analysis of the DNA methylation of maize (Zea mays L.) in response to cold stress based on methylation-sensitive amplified polymorphisms. J Plant Biol 56: 32-38.
- Hao Q, Shang W, Zhang C, Chen H, Chen L, et al. (2016) Identification and comparative analysis of CBS domain-containing proteins in Soybean (Glycine max) and the primary function of GmCBS21 in enhanced tolerance to low nitrogen stress. Int J Mol Sci 17: 620.
- Singh AK, Kumar R, Pareek A, Sopory SK, Singla-Pareek SL (2012) Overexpression of rice CBS domain containing protein improves salinity, oxidative, and heavy metal tolerance in transgenic tobacco. Mol Biotechnol 52: 205-216.
- Ci D, Song Y, Tian M, Zhang D (2015) Methylation of miRNA genes in the response to temperature stress in Populus simonii. Front Plant Sci 6: 921.
- Li J, Huang Q, Sun M, Zhang T, Li H, et al. (2016) Global DNA methylation variations after short- term heat shock treatment in cultured microspores of Brassica napus cv. Topas Sci Rep 6: 38401.
- Liu TK, Li Y, Duan WK, Huang FY, Hou XL (2017) Cold acclimation alters DNA methylation patterns and confers tolerance to heat and increases growth rate in Brassica rapa. J Exp Bot 68: 1213-1224.
- Golldack D, Luking I, Yang O (2011) Plant tolerance to drought and salinity: Stress regulating transcription factors and their functional significance in the cellular transcriptional network. Plant Cell Rep 30: 1383-1391.
- Piao HL, Xuan YH, Park SH, Je BI, Park SJ, et al. (2010) OsCIPK31, a CBL-interacting protein kinase is involved in germination and seedling growth under abiotic stress conditions in rice plants. Mol Cells 30 19-27.
- Kim W, Latrasse D, Servet C, Zhou DX (2013) Arabidopsis histone deacetylase HDA9 regulates flowering time through repression of AGL19. Biochem Biophys Res Commun 432: 394-398.
- Yamaguchi K, Imai K, Akamatsu A, Mihashi M, Hayashi N, et al. (2012) SWAP70 functions as a Rac/Rop guanine nucleotide-exchange factor in rice. Plant J 70: 389-397.
- Zhu N, Cheng S, Liu X, Du H, Dai M, et al. (2015) The R2R3-type MYB gene OsMYB91 has a function in coordinatingplant growth and salt stress tolerance in rice. Plant Sci 236: 146-156.
 Spanish
Spanish  Chinese
Chinese  Russian
Russian  German
German  French
French  Japanese
Japanese  Portuguese
Portuguese  Hindi
Hindi