Research Article, J Biodivers Manage Forestry Vol: 7 Issue: 3
Microbiome Diversity of Endophytic Fungi across Latitudinal Gradients in West Coast Douglas-Fir (Pseudotsuga menziesii) Foliage
Hazel Daniels1, Jed Cappellazzi2 and Jim Kiser3*
1Department of Botany and Plant Pathology, Oregon State University, Corvallis, Oregon, USA
2Department of Wood Science and Engineering, Oregon State University, Corvallis, Oregon, USA
3Department of Forest Engineering, Resources, and Management, Oregon State University, Corvallis, Oregon, USA
*Corresponding Author : Jim Kiser
Department of Forest Engineering, Resources
and Management, Oregon State University, Corvallis, Oregon, USA, Tel: 541-737-
2192; E-mail: jim.kiser@oregonstate.edu
Received: September 04, 2018 Accepted: December 18, 2018 Published: December 24, 2018
Citation: Daniels H, Cappellazzi J, Kiser J (2018) Microbiome Diversity of Endophytic Fungi across Latitudinal Gradients in West Coast Douglas-Fir (Pseudotsuga menziesii) Foliage. J Biodivers Manage Forestry 7:3. doi: 10.4172/2327-4417.1000203
Abstract
Research on forest microbiota lags far behind that of agricultural crops. Understanding of host-pathogen complications is exemplified in defoliation of Douglas-fir dominated forests. Although linked to Phaeocryptopus gaeumannii, etiology of the disease contrasts dramatically with previously known effects. Climate variables and seed source have been suggested as causal, however the number of potential endophytes makes the segregation of a single pathogen for needle cast problematic. A randomized block design was used to assess endophyte populations between seed sources along a latitudinal gradient among a reciprocal provenance study of Douglasfir. Needles were surface sterilized and underwent plate culturing. Isolated fungal colonies were identified through Sanger sequencing and analyzed through BLAST. Differences in seed source and environmental gradient were analyzed by log odds ratio while community relationships were identified through two-way cluster analyses. Linear mixed models were used to evaluate random error associated with seed source data. Forty-six unique isolates from 39 taxa were identified. Infection rates for all needles was 39%, ranging from 5% to 60% among seed sources. Infection rates were higher for non-source needles on three sites. Community structure analyses showed relationships between sample units and endophyte species and strong evidence exists for the influence of seed source type on the relationship between continentality variables and mean average number of endophytes. Ordination demonstrates some evidence for differences between endophyte communities among sites, overcoming any influence provided by seed source type. This study expands the previous known list of Douglas-fir endophytes and outlines fungal community structures based on environmental variables and seed source provenance. This study demonstrated that fungal biodiversity is highly variable in Douglas-fir foliage, and may be influenced by environmental variables, including the provenance of the seed source or the climate at the growing site. Interest in microbiome communities will inevitably lead to advances in management practices.
Keywords: Forest microbiota; Latitudinal gradients; Endophyte
Introduction
Advances in molecular technologies have increased the level of awareness regarding the varied roles of microbiomes (e.g. bacteria and fungi). Assimilated studies of the human microbiota were the among the first areas to address the human body as an integrated ecosystem for example [1]. Research on plant microbiomes has primarily focused on agricultural and nursery crops [2,3]. The complexity of the plant-endophyte relationship has led to the development of novel evolutionary concepts of selection-force interactions on the microbiome collective of both plants and endophytes [4]. However, our understanding of longer-term plant-microbiota systems, such as forests, has been relatively lacking. Implications of this are multifold: foremost, plant pathogenesis has generally been relegated to current visible symptoms but we now understand that pathogenesis can exist as a complex of avirulence/virulence under differing circumstances [5], host-pathogen resistance genes [6], and imbalance in defense mechanisms [7].
The importance of understanding microbiome interactions can be exemplified in current complications with defoliation of Douglas-fir in western Pacific Northwest coastal forests. More recently it has been reported to cause extensive damage to plantations in New England, Europe, and New Zealand [8-10]. In addition to P. gaeumannii, there are other known fungal pathogens associated with Douglas-fir needles. Rhabdocline parkeri has been reported as an asymptomatic fungal infection in Douglas-fir, often described as an avirulent endophyte [11,12]. Rhabdocline pseudotsugae Syd. and R. weirii [13]are both described as endemic but virulent pathogens that cause defoliation in Douglas-fir and bigcone Douglas-fir (Pseudotsuga macrocarpa (Vasey) Mayr.) [14]. Rhizosphaera pseudotsugae was only recently described as a new species, separate from Rhizosphaera kalkhoffii Bubák, both being common in Douglas-fir [15]. Other endophyte species that have been found in Douglas-fir foliage include Bispora sp., Cryptocline abietina Petr.,Geniculosporium sp., Phyllosticta sp., and Xylaria sp.[16]. It is important to note that there isrelatively little information on how the various species known to inhabit Douglas-fir needle tissues interact with other members of their microbial communities.
A preliminary molecular study of Douglas-fir endophytes from Tillamook, Oregon, identified Sydowia polyspora, E. Müll. (anamorph: Hormonema dematioides). In Europe and Canada, S. polyspora is well known for conifer defoliation. It was recently demonstrated in Norway that this fungus is the cause of Current Season Needle Necrosis (CSNN) of Abies sp. and a pathogen of many other conifers [17,18]. In addition to Douglas-fir defoliation, S. polyspora has been implicated as a needle cast pathogen in several studies in Canada and Europe in conifer species including Pinus sp., Abies, sp., Tsuga, sp., Larix,sp., and Picea, sp. The number of known endophytic pathogens, in addition to large numbers of endophytes of indeterminate roles, make the segregation of a single pathogen as the singular cause of needle cast problematic. The importance of this is three-fold. First, if we are to control and/or at least maintain the symptomatic conditions of needle cast, we need to know that we are dealing with the biotic source, and/or biotic complex, that is the underlying foundation for the symptom. A corollary to this is that the coastal forests of Oregon are currently in a more uniform state over many decades of management for single-species even-age stands. This can lead to an imbalance in the pathogen-host relationship where we may actually be seeing a pathogen response that subsequently reestablishes a more naturally diverse stand [19]. In either case, a baseline inventory of the microbiome is necessary. Second, if we are to monitor success in managing disease, especially in terms of restoration, we need to have a baseline to establish some sense of normalcy to the system.
Third, if we are to assess abiotic effects of future conditions (e.g. climate) we require a baseline biota for comparison and analyses. Baseline taxonomic data are crucial to establishing reference points for disease assessment and ecological assessment of change, whether biotic or abiotic in nature.
Methods
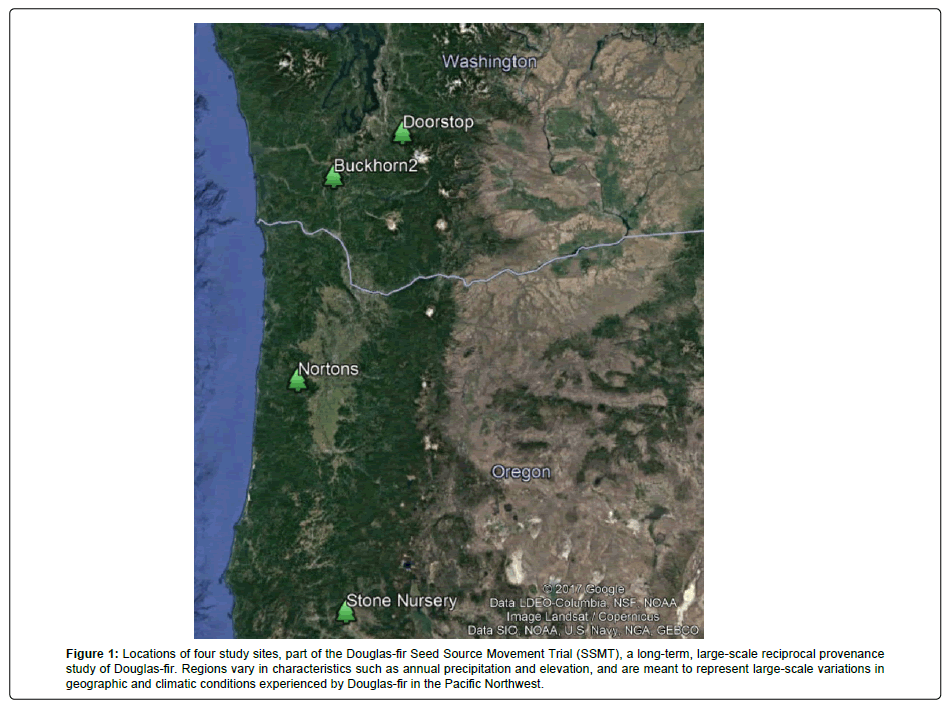
Four study sites were selected along a North-South transect from southern Oregon to central Washington. The study sites are part of an unrelated, but ongoing, study called the Douglas-fir Seed Source Movement Trial (SSMT), a long-term, large-scale reciprocal provenance study of Douglas-fir [20-23]. The four planting sites vary in both elevation (185-860 m) and mean annual precipitation (250-1,575 mm/year) (Figure 1). A nested sampling design was used to collect Douglas-fir branchlets from three different seed sources at each site (Table 2), for a total of twelve seed sources. Within each site, one seed source represents the local population, while the remaining two seed sources represent non-local populations. Non-local seed sources were chosen randomly and without replacement. The second non-local seed source at Doorstop was omitted, as it was a duplicate of the local seed source at Stone Nursery. Therefore, analyses focus on the 11 remaining seed sources (Table 2).
| Site/State | Region | Code | Elevation (m) | MAT* (°C) | MAP (mm) |
|---|---|---|---|---|---|
| Nortons (OR) | Coast North | ORCstN | 185 | 10 | 1198 |
| Stone Nursery (OR) | Cascade Low | ORCasL | 415 | 12 | 250 |
| Buckhorn2 (OR) | Cascade Low | WACasL | 240 | 10 | 1149 |
| Doorstop (WA) | Cascade High | WACasH | 860 | 8 | 1575 |
Table 1: Climate variables associated with seed sources (top) and planting sites (bottom) of the Seed Source Movement Trial (SSMT), a long-term, large-scale reciprocal provenance study of Douglas-fir.
| Planting site | Local seed source | Non-local seed sources | |
|---|---|---|---|
| Buckhorn2 | WACasL | WACst | ORCasL |
| Doorstop | WACasH | CAKla | (ORSisL—OMITTED) |
| Nortons | ORCstN | ORCstS | ORCasH |
| Stone Nursery | ORSisL | CACst | CASierra |
Table 2: Seed sources sampled at each site. Planting sites are part of the Seed Source Movement Trial (SSMT), a long-term, large-scale reciprocal provenance study of Douglas-fir.
Figure 1: Locations of four study sites, part of the Douglas-fir Seed Source Movement Trial (SSMT), a long-term, large-scale reciprocal provenance study of Douglas-fir. Regions vary in characteristics such as annual precipitation and elevation, and are meant to represent large-scale variations in geographic and climatic conditions experienced by Douglas-fir in the Pacific Northwest.
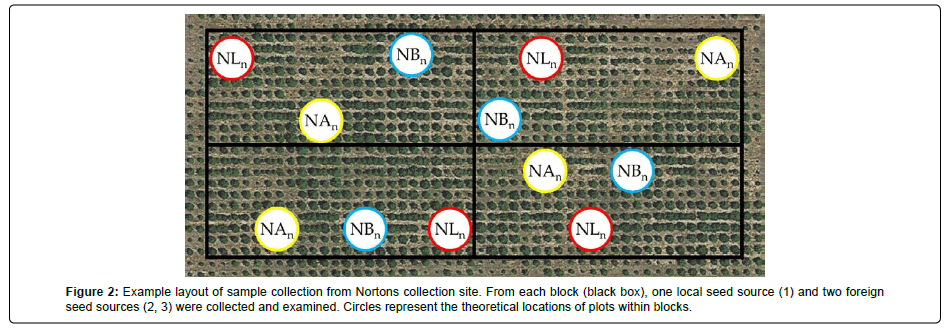
Two to four blocks were sampled at each site, depending on the availability of living samples (Figure 2). For several seed sources, at least one block contained no living trees of that source and could not be sampled. Within each section, one arbitrarily chosen tree and one branchlet were sampled. The branchlet position on each tree varied both in vertical position (high, medium, or low) and cardinal direction (N, S, E, W). Vertical and directional variations were based on branchlet availability within reach. Branchlets were stored at -20°C prior to analysis. Thirty-seven total branchlets were collected.
Traditional plate culturing
Needles were removed from each branchlet and surface-sterilized in 2% bleach for 5 minutes, then rinsed in deionized water and transferred to a sterile 15 mL collection tube [3]. Needles were randomly chosen from the collection tube for sterile plating, cut into 5 or more pieces, and placed onto 1.5% potato dextrose agar (PDA) [Difco™ (Becton, Dickinson and Company, Sparks, MD, USA)]. Each plate was evenly divided into three observation units, with one sectioned needle placed in each unit. For each seed source, 5 PDA plates were used, for a total of 15 needles from each seed source. Plates were visually observed daily for evidence of fungal growth. Newly observed growth was immediately transferred to a 1.5% PDA plate to isolate pure cultures. After at least fourteen days of culturing, isolates were examined and grouped based on colony morphology. One representative of each morphogroup was chosen for sequencing; however, a conservative approach was applied, where morphological uncertainties were sequenced to prevent species omission. Pure fungal tissue was collected from potato dextrose broth incubations for molecular analysis. Surface sterilization effectiveness was tested using a sub-sample from stored collection tubes and methods described in [3]. No plates exhibited growth originating from imprinted areas, suggesting surface sterilization was effective.
DNA extraction
Isolate DNA was extracted using a DNeasy Plant Mini Kit® (QIAgen, Valencia, CA, USA), according to standard protocol. Fourteen isolates could not be extracted due to slow or absent growth in liquid media. DNA was extracted from these isolates using a modified cetyl trimethylammonium bromide (CTAB) DNA extraction protocol [24].
Isolate extracts underwent DNA amplification via polymerase chain reaction (PCR) according to Cappellazzi et al. [1]. Following PCR, amplicons were checked on a 1.5% agarose gel stained with the Nucleic Acid Staining Solution RedSafe™ (iNtRON Biotechnology, South Korea) for a single, target fungal band. PCR product was purified with Exo-Sap™ PCR product cleanup kit (Affymetrix, Santa Clara, CA, USA) following manufacturer protocol. Purified product was submitted to the Center for Genome Research and Biocomputing, where Sanger sequencing was performed with an ABI 3730 capillary sequence machine (Life Technologies, Grand Island, NY, USA). Completed sequences were assembled into contigs using Sequencher 4.9 and fungal identities were determined using the Basic Local Alignment Search Tool (BLAST) to identify species.
Data analysis
Data were analyzed for community structure within needles, seed sources, and sites using PC-ORD Version 7.0 [2]. Sample units (needles) were only included in the data set if they had successfully sequenced endophytes. Unamplified sequences which could not be identified via BLAST were removed from the analysis, but needles with multiple isolates were retained so long as at least one isolate was identified. A number of the unidentified isolates were visually identified as bacterial, which would make identification and sequencing with fungal ITS regions unlikely. From 555 needles tested, 155 needles fulfilled the above criterion.
Two data sets were created to analyze community data. The first data set, “Endophytes by Needle (EN)”, included a species matrix (155 sample units x 46 species), which contained counts of individual endophytes on each needle. An environmental matrix (155 sample units x 29 environmental variables) contained measurements of environmental variables collected in 2013, and experimental design variables (seed source) (Table 3). The second data set, “Endophytes by Seed Source (ESS)”, was created from the grouped sample units, and included all endophytes from all needles of a seed source. Since each seed source was only collected at one site, they are within-site groups. The species matrix (11 seed sources x 46 species) contained presence/absence information for endophyte species. The environmental matrix (11 seed sources x 29 environmental variables) was the same as that of the EN data set (Table 3). Since ESS data were combined within-site, the same environmental variables could be used. Data were not relativized or otherwise transformed.
| Variable | Definition |
|---|---|
| CMD_wt | Winter (Dec – Feb) Hargreaves climatic moisture deficit (mm) |
| CMD_sp | Spring (Mar – May) Hargreaves climatic moisture deficit (mm) |
| CMD_sm | Summer (Jun – Aug) Hargreaves climatic moisture deficit (mm) |
| CMD_at | Autumn (Sept – Nov) Hargreaves climatic moisture deficit (mm) |
| RH_wt | Winter relative humidity (%) |
| RH_sp | Spring relative humidity (%) |
| RH_sm | Summer relative humidity (%) |
| RH_at | Autumn relative humidity (%) |
| RH_avg | Average relative humidity (%) |
| MWMT | Mean warmest month temperature (°C) |
| MCMT | Mean coldest month temperature (°C) |
| TD | Temperature difference between MWMT and MCMT; continentality (°C) |
| MAP | Mean annual precipitation (mm) |
| MSP | Mean annual summer (May – Sept) precipitation (mm) |
| NFFD | Number of frost-free days |
| FFP | Frost-free period |
| PAS | Precipitation as snow (mm) between Aug in previous year and Jul in current year |
| Eref | Hargreaves reference evaporation (mm) |
| CMD | Hargreaves climatic moisture deficit (mm) |
| Tave_wt | Winter mean temperature (°C) |
| Tmin_wt | Winter mean minimum temperature (°C) |
| DD_0 | Degree-days below 0 °C, chilling degree-days |
| DD5 | Degree days above 5 °C, growing degree-days |
| DD_18 | Degree-days below 18 °C. heating degree-days |
| DD18 | Degree-days above 18 °C, cooling degree-days |
Table 3: Environmental and experimental variables utilized in community data analysis.
In order to visually assess endophyte community relationships, a two-way cluster analysis was performed on both EN and ESS data sets. The analyses used Euclidean distance, Ward’s clustering method, and relativization by variable maximum. Multi-Response Permutation Procedures (MRPP) were performed on EN and ESS datasets, using site as the grouping identifier and Sørensen distance [25]. Additionally, non-metric multidimensional scaling (NMS) was used with the EN data set to examine variation among sample units (needles) with respect to endophytes. NMS ordinations use ranked-distances to plot similar sample units closer together. The distance between points/sample units widens with increasing dissimilarity in endophyte populations [26,27]. NMS is well suited to data for which variables do not have linear relationships, unlike other ordination systems [25]. For analysis of the EN dataset, the NMS ordination was performed in autopilot mode, using medium thoroughness and Euclidean distance, in PC-ORD Version 7.0 [28]. Ties, where ranked distances are the same in two or more sample units, were not penalized (Kruskal’s primary approach) [27]. Finally, outlier analysis was performed on both EN and ESS data sets using Euclidean distance measure.
Odds ratios were calculated between sites and between local and non-local seed sources within sites to describe the binary responses for the data collected [29]. Mean average endophyte number by seed source type (local, non-local) at various environmental variables (Table 3) was also calculated. No comparisons were made across sites in the calculation of mean average endophyte number, since the nested study design precludes such comparisons. Since non-local seed sources can come from drastically different conditions than local seed sources [20,30,31], it is expected that the mean average endophyte number will differ between local and non-local seed sources across multiple variables.
Evaluation of differences in mean average number of endophytes between local and non-local seed sources within continuous variables were included in this study (ENV-VAR). Linear mixed models were designed to account for the random error associated with each observation (average number of endophytes per needle per seed source type). Analyses were performed without adjustment to the α-level, since only two comparisons were performed within sites (α=0.95). All analyses were performed using R version 3.4.1 [3], and all R graphics were created with the ggplot2 package [32]. The following statistical model was used to describe the linear mixed model for the data collected from the design outlined above:
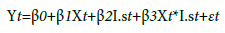
Where:
Yt is the mean average number of endophytes from i seed source type from j site; i=local, non-local; j=Stone, Buckhorn2, Nortons, Doorstop,
β0 is the mean average number of endophytes in the local seed sources when ENV-VAR is 0,
β1 is the incremental effect of ENV-VAR on the mean average number of endophytes in the
local seed source,
β2 is the incremental effect of seed source type on the mean average number of endophytes
when ENV-VAR is 0,
β3 is the incremental effect of the interaction between average number of endophytes and seed
source type on the number of endophytes at a site,
Xt is a continuous variable representing the average number of endophytes from the jth site,
I.s is 1 when the seed source is non-local and 0 otherwise, εt is the random error associated with the tth observation, εt ~ N(0, σ2) and εt and εt’ are independent.
β values were combined to create various estimates of means. For a local seed source, β0, β1, and εt were summed to get an estimate of the mean average number of endophytes. For non-local seed sources, the estimate of the mean average number of endophytes is the product of: (β0+β2)+(β1+β3)Xt+εt. It should be noted that, because each site has a unique precipitation level with no overlap between sites, these two factors are confounded and interchangeable, which prevents the comparison of average mean numbers of endophytes across sites.
Results
Of the 555 Douglas-fir needles sampled, 318 isolates were cultured from 215 needles. Eighty-nine isolates were sorted into 54 morphogroups, of which 34 were successfully identified using BLAST lookup. There were 126 ungrouped isolates, of which 96 were successfully identified. The remaining 20 morpho groups and 30 ungrouped isolates remain unidentified due to lack of amplification during PCR, or poor-quality sequencing resulting in unsuccessful identification. In total, 46 unique isolates from 39 taxa were identified from needles (Table 4).
| Species | Frequency | Accession Number(s) |
|---|---|---|
| Anthostomella conorum1 | 6 | EU552099, KT149745 |
| Anthostomella pinea | 1 | KJ406991 |
| Apodus deciduus | 1 | AY681199 |
| Aposphaeria corallinolutea | 1 | KY554202 |
| Aureobasidium pullulans | 4 | LC277152, JX188099 |
| Aureobasidium sp. | 1 | HF674760 |
| Cladosporium perangustum | 1 | MF303712 |
| Claussenomyces sp.1 | 2 | KT264343 |
| Clypeosphaeria mamillana2 | 4 | KT949898 |
| Clypeosphaeria sp. | 1 | JQ341099 |
| Coniochaeta hoffmannii | 1 | KX869937 |
| Crustomyces subabruptus1 | 24 | KP814558 |
| Cryptococcus sp. | 1 | KM216339 |
| Cryptostroma corticale2 | 3 | KR870994 |
| Diaporthe sp.1 | 2 | LC041016 |
| Dothideomycetes sp. | 4 | KP990991 |
| Geopyxis rehmii | 2 | KU932461 |
| Graphostroma sp.1,2 | 13 | EU715682 |
| Helicoon fuscosporum | 2 | EF029203 |
| Hormonemia sp. | 2 | AF013225 |
| Hypoxylon rubiginosum2 | 21 | AY787708 |
| Lecythophora sp.1,2 | 7 | KX096678, GU062252, AY219880 |
| Melanomma pulvis-pyrius | 2 | KY189979 |
| Nemania serpens | 1 | KU141386 |
| Nemania sp. | 1 | HM123573 |
| Ophiognomonia alni-viridis | 1 | JF514848 |
| Penicillium glabrum1,2 | 18 | KU847873, KY318471, KX099660, KX609402, KU847870 |
| Perusta inaequalis1 | 2 | NR_144958 |
| Pezizomycetes sp.1 | 3 | KX909069, KJ508333 |
| Phaeomoniella zymoides | 2 | GQ154600 |
| Phomatospora biseriata | 1 | KX549454 |
| Podospora sp. | 1 | AM262361 |
| Preussia bipartis2 | 3 | GQ203774 |
| Rhabdocline parkeri1 | 8 | AF462428, AF462427 |
| Rosellinia quercina | 3 | AB017661 |
| Rosellinia sp.1 | 3 | KT264658 |
| Sordariales sp.1 | 2 | FN548158 |
| Sordariomycetes sp.1,2 | 9 | KT264524, KX611007, KX611549, JQ759589, GQ153043, KP992078, GQ153206 |
| Sporormiaceae sp. | 1 | KX611024 |
| Sydowia polyspora | 1 | KP152486 |
| Talaromyces ruber1 | 2 | JX965239 |
| Taphrina communis1 | 13 | AF492088 |
| Taphrina veronaerambellii | 1 | NR_111148 |
| Wallemia sebi | 1 | KX911858 |
| Xylaria hypoxylon1,2 | 35 | KY204024, KX096696, GU300096 |
| Xylaria sp.2 | 3 | AB465207 |
| Unidentified | 97 |
Table 4: Taxa and frequency of endophyte species cultured and sequenced from 185 Douglas-fir needles from four sites in Oregon and Washington. The number of endophytes successfully sequenced totals 221 out of 318 isolates.
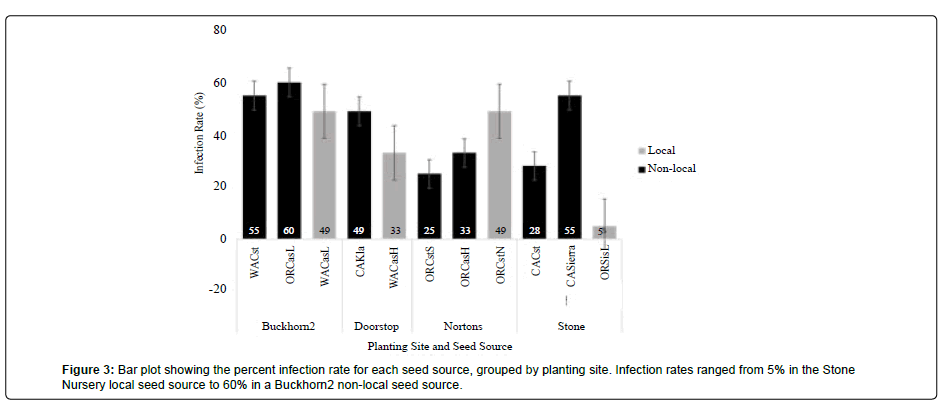
Rates of infection for all needles was 39%, with infection ranging from 5% to 60% among seed sources (Table 5, Figure 3). At Buckhorn2, Doorstop, and Stone Nursery, infection rates of needles from non-local seed sources were higher than those of needles from local seed sources.
| Site name | Type | Seed source | Isolates | Needles cultured | % Infection rate |
|---|---|---|---|---|---|
| Buckhorn2 | Non-local | WACst | 33 | 60 | 55% |
| Buckhorn2 | Non-local | ORCasL | 18 | 30 | 60% |
| Buckhorn2 | Local | WACasL | 22 | 45 | 49% |
| Doorstop | Non-local | CAKla | 22 | 45 | 49% |
| Doorstop | Local | WACasH | 15 | 45 | 33% |
| Nortons | Non-local | ORCstS | 15 | 60 | 25% |
| Nortons | Non-local | ORCasH | 15 | 45 | 33% |
| Nortons | Local | ORCstN | 22 | 45 | 49% |
| Stone | Non-local | CACst | 17 | 60 | 28% |
| Stone | Non-local | CASierra | 33 | 60 | 55% |
| Stone | Local | ORSisL | 3 | 60 | 5% |
| Total: | 215 | 555 | 39% |
Table 5: Endophyte infection rates for each seed source sampled at four sites in Oregon and Washington.
The odds of infection of a non-local needle at Buckhorn2 was 1.4 to 1 (Fisher’s exact test, one-sided p-value=0.25). Non-local needles at Doorstop were 1.9 times as likely as local needles to have an infection (p=0.099). At Nortons, the odds of infection of a local needle were 2.4 times higher than that of non-local needles (p=0.01). The odds of infection at Stone Nursery was 13.6 times more likely in non-local than local needles (p<0.0001). Based on our methods, the odds of finding an endophyte infection on a non-local seed source was 1.6 times higher overall (p=0.007).
Confidence intervals for these odds ratios, and between site comparisons of infection odds ratios can be seen in Table 6. Sixteen unique taxa were found at more than one site (Table 6). Ten taxa were isolated within needles of both local and non-local seed sources within sites. Isolates of Graphostroma sp., Lecythophora sp., Penicillium glabrum, Sordariomycetes sp., and Xylaria hypoxylon were found in local and non-local needles at multiple sites. Penicillium glabrum and Sordariomycetes sp. were found at all four sites, with P. glabrum isolates at three different sites sharing the same accession number. Similarly, X. hypoxylon were isolated from needles at both Buckhorn2 and Nortons. Out of 32 isolates, 28 shared the same accession number. Examination of community structure via two-way cluster analyses for both ESS and EN revealed relationships between sample units and endophyte species. The ESS cluster analysis shows a few natural groupings in seed sources, such as those between ORSisL, CACst, and WACasH. There are also groupings on the species side, where some species only occur alongside specific other species. There was 8% chaining among rows in the ESS cluster but only 3% chaining among columns, indicating strong clustering between seed sources and between endophytes. The EN analysis was less robust, with 8% chaining among rows, but 81% chaining among columns. Because most needles have only one isolated endophyte, there are no endophyte communities at the needle level. Analysis of species richness (α diversity) by seed source (ESS data set) ranged from 1.1 for ORSisL (Stone Nursery local seed source) to 2.6 for WACst (Buckhorn2 non-local seed source), with an average of 2.1 for all sites (Table 7).
| Confidence Interval (95%) | ||||
|---|---|---|---|---|
| Comparison | Odds ratio | Lower limit | Upper limit | p-value |
| Buckhorn2 Non-Local:Local | 1.3671 | 0.6668 | 2.803 | 0.2507 |
| Doorstop Non-Local:Local | 1.913 | 0.8161 | 4.4845 | 0.0992 |
| Nortons Local:Non-Local | 2.3913 | 1.1619 | 4.9214 | 0.0143 |
| Stone Nursery Non-Local:Local | 13.5714 | 4.0211 | 45.804 | <0.0001 |
| Buckhorn2:Doorstop | 1.6866 | 0.9835 | 2.8923 | 0.0382 |
| Buckhorn2:Nortons | 2.219 | 1.377 | 3.5758 | 0.0007 |
| Buckhorn2:Stone Nursery | 2.8214 | 1.7701 | 4.4969 | <0.0001 |
| Doorstop:Nortons | 1.3157 | 0.7683 | 2.2531 | 0.1940 |
| Doorstop:Stone Nursery | 1.6728 | 0.9864 | 2.8371 | 0.0382 |
| Nortons:Stone Nursery | 1.2715 | 0.799 | 2.0233 | 0.1852 |
Table 6: Odds ratios comparing incidence of infection within and between four sites in Oregon and Washington.
| Diversity measure | ||||
|---|---|---|---|---|
| Data set | N | α | β | γ |
| Endophytes by Needles (EN) | 155 | 0.2 | 35.2 | 7 |
| Endophytes by Seed Source (ESS) | 11 | 2.1 | 4.4 | 9 |
Table 7: Alpha, beta, and gamma species diversity of endophytes in Douglas-fir needles from EN and ESS data sets.
Analysis of endophytes by needle (data set EN) had a low average α diversity (0.2) due to the large number of needles with only a single isolated endophyte species. Between-site heterogeneity (β diversity) and species richness over a range of habitats (γ diversity) is also shown in Table 7. MRPP tested the null hypothesis of no difference between groups. For both the EN and ESS data sets, MRPP provided strong evidence to reject the null hypothesis (Tables 8 and 9). In all cases, observed δ (weighted mean within-group distance) is smaller than expected by chance, and the probability of a δ equal to or less than the observed value is very small. In addition, A, the chance-corrected within-group agreement, is between 0 and 1, indicating there is more heterogeneity within groups than expected by chance. Pairwise comparisons were also performed, comparing each site to the others. The p-values for these pairwise comparisons were not corrected for multiple comparisons (i.e. Bonferroni correction). Ten of the twelve pairwise comparisons have statistically significant p-values.
| Groups | Observed δ | Expected δ | T | p-value | A |
|---|---|---|---|---|---|
| Sørensen | 0.66 | 0.80 | -3.30 | 0.003 | 0.18 |
| Multiple Comparisons (Sørensen) | |||||
| Buckhorn2 vs. Doorstop | -1.06 | NaN | 0.07 | ||
| Buckhorn2 vs. Nortons | -1.96 | 0.041 | 0.14 | ||
| Buckhorn2 vs. Stone Nursery | -1.88 | 0.037 | 0.11 | ||
| Doorstop vs. Nortons | -1.90 | 0.000 | 0.21 | ||
| Doorstop vs. Stone Nursery | -0.32 | NaN | 0.03 | ||
| Nortons vs. Stone Nursery | -2.51 | 0.025 | 0.23 |
Table 8: Multi-response Permutation Procedures results for ESS data set. Note that multiple comparison p-values were not corrected for multiple comparisons (i.e. Bonferroni).
| Groups | Observed δ | Expected δ | T | p-value | A |
|---|---|---|---|---|---|
| Sørensen | 0.85 | 0.93 | -20.31 | 0.000 | 0.08 |
| Multiple Comparisons (Sørensen) | |||||
| Buckhorn2 vs. Doorstop | -10.48 | 0.000 | 0.05 | ||
| Buckhorn2 vs. Nortons | -10.34 | 0.000 | 0.04 | ||
| Buckhorn2 vs. Stone Nursery | -8.72 | 0.000 | 0.04 | ||
| Doorstop vs. Nortons | -15.26 | 0.000 | 0.07 | ||
| Doorstop vs. Stone Nursery | -6.88 | 0.000 | 0.05 | ||
| Nortons vs. Stone Nursery | -10.62 | 0.000 | 0.07 |
Table 9: Multi-response Permutation Procedures results for EN data set. Note that multiple comparison p-values were not corrected for multiple comparisons (i.e. Bonferroni).
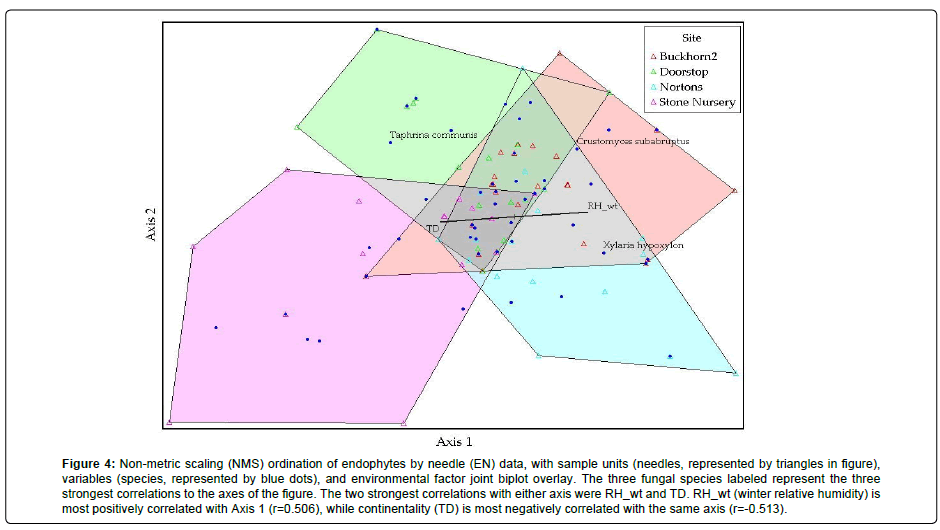
NMS results for the EN data set converged on a stable two-dimensional solution (Table 10). The final stress of the two-dimensional solution was 9.24, with a final instability of 0.0000 from 94 iterations. This solution also had a chance-corrected final improvement value of 0.77 (1 being a perfect fit). Dimensionality was assessed, and the two-dimensional solution offered the greatest improvement, with all subsequent solutions offering insufficient improvement for use in analysis. Cumulatively, 56.25% of variation can be attributed to the two axes chosen by the model. Endophyte species with the strongest correlations were Xylaria hypoxylon (Axis 1 r=0.577), Taphrina communis (Axis 2 r=0.352), and Crustomyces subabruptus. (Axis 2 r=0.350). In the NMS 2D ordination graph, winter relative humidity (RH_wt) was most positively correlated with Axis 1 (r=0.506), while continentality (TD) was most negatively correlated with that same axis (r=-0.513) (Figure 4). Longitude (r=0.435) and autumn relative humidity (r=0.426) had the strongest positive correlations with Axis 2, and mean annual temperature (MAT) had the strongest negative correlation with Axis 2 (r=-0.428). PC-ORD was not able to find a useful NMS ordination for the ESS data set, due to weakly structured data (Table 11). Outlier analysis did not find any outliers in either data set.
| Stress in real data | Stress in randomized data | ||||||
|---|---|---|---|---|---|---|---|
| Axes | Minimum | Mean | Maximum | Minimum | Mean | Maximum | p-value |
| 1 | 17.59 | 26.87 | 40.44 | 29.76 | 32.51 | 35.51 | 0.0196 |
| 2 | 8.94 | 9.86 | 11.00 | 14.05 | 16.21 | 20.33 | 0.0196 |
| 3 | 5.28 | 5.82 | 6.51 | 8.43 | 9.34 | 10.72 | 0.0196 |
| 4 | 3.54 | 3.87 | 4.40 | 5.44 | 6.19 | 7.32 | 0.0196 |
Table 10: Stress in relation to dimensionality (number of axes) comparing 50 runs on the real data with 50 runs on randomized data from the EN data set. The p-value is the proportion of randomized runs with stress less than or equal to the observed stress. The two-dimensional solution is highlighted for use in further analysis.
| Stress in real data | Stress in randomized data | ||||||
|---|---|---|---|---|---|---|---|
| Axes | Minimum | Mean | Maximum | Minimum | Mean | Maximum | p-value |
| 1 | 27.92 | 39.74 | 52.21 | 19.51 | 35.22 | 51.64 | 0.294 |
| 2 | 12.99 | 16.32 | 21.79 | 4.86 | 13.48 | 24.78 | 0.451 |
| 3 | 5.32 | 6.73 | 9.17 | 2.07 | 2.07 | 9.60 | 0.412 |
| 4 | 0.55 | 1.39 | 4.85 | 0.01 | 1.91 | 4.60 | 0.196 |
Table 11: Stress in relation to dimensionality (number of axes) comparing 50 runs on the real data with 50 runs on randomized data from the ESS data set. The p-value is the proportion of randomized runs with stress less than or equal to the observed stress. No useful NMS ordination was found from this data.
Figure 4: Non-metric scaling (NMS) ordination of endophytes by needle (EN) data, with sample units (needles, represented by triangles in figure), variables (species, represented by blue dots), and environmental factor joint biplot overlay. The three fungal species labeled represent the three strongest correlations to the axes of the figure. The two strongest correlations with either axis were RH_wt and TD. RH_wt (winter relative humidity) is most positively correlated with Axis 1 (r=0.506), while continentality (TD) is most negatively correlated with the same axis (r=-0.513).
The environmental variables used in statistical analysis were based on the outcome of the NMS ordination as well as previous research (4,5). Continentality (TD) and winter relative humidity (RH_wt) were chosen at the two most relevant variables from the NMS ordination, and mean coldest month temperature (MCMT) was chosen based on Stone and Mater [5]. Each variable was analyzed separately with relation to seed source type. Model assumptions were tested and confirmed for each interaction analysis. To address whether the relationship of mean number of fungal endophytes and each environmental variable (TD, RH_wt, MCMT) differed between local and non-local seed sources, the total number of endophytes for a seed source type was averaged without regard to the number of needles sampled. This was to find a number of endophytes rather than a number of endophytes per needle. All endophyte counts from non-local seed sources at a site were averaged to create a mean average number of endophytes per non-local seed source. Using the ANOVA function in R, analyses were performed on the mean average number of endophytes (Tables 11-13). There was strong evidence for the influence of seed source type on the relationship between continentality (Table 12) and mean average number of endophytes but not winter relative humidity (Table 13) or mean coldest month temperature (Table 14). Estimated variances (σ2) were 18.29 (TD), 20.48 (RH_wt), and 32.02 (MCMT). Differences between mean average number of endophytes and 95% confidence intervals were constructed based on the model outputs (Tables 11-13). The incremental effect of continentality (TD) on mean average number of endophytes for a local seed source was estimated to be 2.24 endophytes fewer than the default (local seed source with TD of 0) (95% CI: [-4.018, -0.461]). The incremental effect of non-local seed source on mean average number of endophytes at a site with TD of 0 was estimated to be 45.76 endophytes fewer than the default (95% CI: [-93.172, -1.643]). The incremental effect of the interaction between TD and non-locaseed source was estimated to be 2.81 endophytes more than the control (95% CI: [0.295, 5.326]. The incremental effect of winter relative humidity (RH_wt) on mean average number of endophytes for a local seed source was estimated to be 2.55 endophytes more than the default (local seed source with RH_wt of 0) (95% CI: [0.415, 4.678]). The incremental effect of non-local seed source on mean average number of endophytes at a site with RH_wt of 0 was estimated to be 221.75 endophytes more than the default (95% CI: [-3.731, 447.235]). The incremental effect of the interaction between RH_wt and non-local seed source was estimated to be 2.88 endophytes fewer than the control (95% CI: [-5.895, 0.133].
| Df | Sum Sq | Mean Sq | F-value | p-value | |
|---|---|---|---|---|---|
| TD | 1 | 61.972 | 61.972 | 3.388 | 0.139 |
| SSType | 1 | 81.281 | 81.281 | 4.444 | 0.103 |
| TD : SSType | 1 | 176.058 | 176.058 | 9.626 | 0.036 |
| Residuals | 4 | 73.157 | 18.289 |
Table 12: ANOVA output for interaction of continentality (TD) and seed source type on mean average number of endophytes within each planting site. Response: mean average number of endophytes per seed source.
| Df | Sum Sq | Mean Sq | F-value | p-value | |
|---|---|---|---|---|---|
| RH_wt | 1 | 85.033 | 85.033 | 4.153 | 0.111 |
| SSType | 1 | 81.281 | 81.281 | 3.969 | 0.117 |
| RH_wt : SSType | 1 | 144.245 | 144.245 | 7.044 | 0.057 |
| Residuals | 4 | 81.910 | 20.478 |
Table 13: ANOVA output for interaction of winter relative humidity (RH_wt) and seed source type on mean average number of endophytes within each planting site. Response: mean average number of endophytes per needle.
| Df | Sum Sq | Mean Sq | F-value | p-value | |
|---|---|---|---|---|---|
| MCMT | 1 | 42.326 | 42.326 | 1.321 | 0.314 |
| SSType | 1 | 81.281 | 81.281 | 2.538 | 0.186 |
| MCMT : SType | 1 | 140.763 | 140.763 | 4.396 | 0.104 |
| Residuals | 4 | 128.098 | 32.025 |
Table 14: ANOVA output for interaction of mean coldest month temperature (MCMT) and seed source type on mean average number of endophytes within each planting site. Response: mean average number of endophytes per needle.
The incremental effect of mean coldest month temperature (MCMT) on mean average number of endophytes for a local seed source was estimated to be 2.23 endophytes fewer than the default (local seed source with MCMT of 0) (95% CI: [-4.935, 0.468]). The incremental effect of non-local seed source on mean average number of endophytes at a site with MCMT of 0 was estimated to be 49.52 endophytes fewer than the default (95% CI: [-124.369, 25.331]). The incremental effect of the interaction between MCMT and non-local seed source was estimated to be 2.88 endophytes more than the control (95% CI: [-0.936, 6.705].
Discussion
The NMS ordination demonstrates some evidence that there are differences between endophyte communities among sites, and that those differences overcome any influence provided by seed source type. The cluster analyses indicate natural grouping in the endophytes by seed source (ESS) data beyond that inherent in the test itself. The relatedness of Stone Nursery and Doorstop seed sources is one example of that. However, the clustering overall is grouped by site, with each site’s three seed sources (one local and two non-local) clustered closer together. This is further evidence that sites themselves are more influential on endophyte communities than seed source. The cluster analysis of the endophytes by needle (EN) data set demonstrated how the high beta diversity makes the data difficult to analyze in its current state. Multi-Response Permutation Procedures (MRPP) demonstrated statistically significant differences between groups, which are reflected in the pairwise comparisons. The NMS ordination graph shows distinct separations between all sites, though there is also significant overlap. There appears to be at least some influence of continentality and winter relative humidity on endophyte populations. These endophytes all share the same host species, Douglas-fir, so it should not be surprising to see many of the same species within needles at different sites. What was surprising to observe were the relatively weak correlations between specifically measured environmental factors and endophyte populations according to the NMS ordination.Statistical analysis provides evidence that the relationship of mean number of fungal endophytes and certain environmental variables differ between local and non-local seed sources for these 11 seed sources at the four sites outlined above (Table 2). This is separate from the ordination results, as they focus on the species and their community structures, while the statistical analysis focuses instead on counts and frequencies of endophytes regardless of species.
According to the model, continentality (TD) has little influence on the mean average number of endophytes alone, yet the interaction between TD and seed source type has a statistically significant influence on the explanatory variable. Analysis revealed the possibility that non-local seed source, regardless of environmental variable, leads to higher mean average number of endophytes, although the 95% confidence interval for the difference in mean average number of endophytes contains 0 in all cases. Based on that confidence interval, non-local seed source could also lead to a biologically significant reduction in mean average number of endophytes, as the confidence interval includes both the positive and negative 20% threshold.
Conclusion
This is the first study to identify endophytes of coastal Oregon Douglas-fir through molecular methods. It has expanded the previously known list of Douglas-fir endophytes put forth by Carroll and Carroll and outlines fungal community structures based on environmental variables and seed source provenance within the limitations of the study [16]. Further, it provides evidence that the microbial community of Douglas-fir is a topic deserving of further attention, as it may have tangible effects on plantation management and assisted migration of Douglas-fir seedlings. This study demonstrated that fungal biodiversity is high in Douglas-fir foliage, and may be influenced by environmental variables, including the provenance of the seed source or the climate at the growing site. Endophytes are important factors in the management of plants and forest trees [33], so the growing interest in microbiome communities will inevitably lead to advances in best management practices, potentially including steps such as inoculation with local beneficial endophytes to stave off infection by critical mass as seen with P. gaeumannii. The fine-grain structure of endophyte communities in Douglas-fir, especially those in relation to P.gaeumannii, and their relation to environmental conditions should be more deeply addressed in future studies.
Acknowledgement
The authors gratefully acknowledge Amy Klocko, Steve Strauss, Anna Magnusen, and Michael Freitag for the use of workspace and hardware, Lisa Ganio and Ariel Muldoon for statistical advising, and Connie Harrington for assistance with study design.
References
- Huttenhower C, Gevers D, Knight R, Abubucker S, Badger JH, et al. (2012) Structure, function and diversity of the healthy human microbiome. Nature 486: 207-214.
- Saunders DGO (2014) Hitchhiker's guide to multi-dimensional plant pathology. New Phytologist205: 1028-1033.
- Schulz B, Wanke U, Draeger S, Aust HJ (1993) Endophytes from herbaceous plants and shrubs: effectiveness of surface sterilization methods. Mycol Res 97: 1447-1450.
- Hardoim PR, Van Overbeek LS, Berg G, Pirttilä AM, Compant S, et al. (2015) The hidden world within plants: Ecological and evolutionary considerations for defining functioning of microbial endophytes. Microbiol Mol Biol Rev 79: 293-320.
- Junker C, Draeger S, Schulz B (2012) A fine line-endophytes or pathogens in Arabidopsis thaliana. Fungal Ecol 5: 657-662.
- Schafer W (1994) Molecular mechanisms of fungal pathogenicity to plants. Annu Rev Phytopathol32: 461-477.
- Schulz B, Boyle C (2005) The endophytic continuum. Mycol Res 109: 661-686.
- Hood IA, Kershaw DJ (1975) Distribution and infection period of Phaeocryptopusgaeumannii in New Zealand. NZJ For Sci 5: 201-208.
- Lakomy P, Iwanczuk M (2010) Phaeocryptopus gaeumannii in Douglas-fir stands in Smolarz forest district. Phytopathologia 58: 43-52.
- Merrill W, Longenecker J (1973) Swiss needle cast on Douglas-fir in Pennsylvania. Plant Disease Reporter57: 984.
- Sherwood-Pike M, Stone JK, Carroll GC (1986) Rhabdocline parkeri, a ubiquitous foliar endophyte of Douglas-fir. Can J Bot 64: 1849-1855.
- Stone JK (1987) Initiation and development of latent infections by Rhabdocline parkeri on Douglas-fir. Can J Bot 65: 2614-2621.
- Parker AK, Reid J (1969) The genus Rhabdocline Syd. Can J Bot 47: 1533-1545.
- Chastagner GA (2001) Susceptibility of intermountain douglas-fir to rhabdocline needle cast when grown in the Pacific Northwest. Plant Health Prog.
- Butin H, Kehr R (2000) Rhizosphaera pseudotsugae sp. nov. and related species. Mycol Res104: 1012-1016.
- Carroll GC, Carroll FE (1978) Studies on the incidence of coniferous needle endophytes in the Pacific Northwest. Can J Bot 56: 3034-3043.
- Smerlis E (1970) Notes on Sydowia polyspora. Can J Bot 48: 1613-1615.
- Talgø V, Chastagner G, Thomsen IM, Cech T, Riley K, et al.(2010) Sydowia polyspora associated with current season needle necrosis (CSNN) on true fir (Abies spp.). Fungal Biol 114: 545-554.
- Manion PD (1991) Tree Disease Concepts. (2nd edtn), Prentice-Hall, New Jersey, USA.
- Bansal S, Harrington CA, Gould PJ, St. Clair JB (2015) Climate-related genetic variation indrought-resistance of Douglas-fir (Pseudotsuga menziesii). Glob Chang Biol 21: 947-958.
- Ford KR, Harrington CA, Bansal S, Gould PJ, St. Clair JB (2016) Will changes in phenology track climate change? A study of growth initiation timing in coast Douglas-fir. Glob Chang Biol 22: 3712-3723.
- Gould PJ, Harrington CA, St.Clair JB (2010) Incorporating genetic variation into a model of budburst phenology of coast Douglas-fir (Pseudotsuga menziesii var. menziesii). Can J For Res 41: 139-150.
- Gould PJ, Harrington CA, Clair JBS (2012) Growth phenology of coast Douglas-fir seed sources planted in diverse environments. Tree Physiol 32: 1482-1496.
- M. Gardes, T. Bruns (1996) Species Diagnostics Protocols: PCR and Other Nucleic Acid Methods. Humana Press, Totowa, NJ, USA.
- McCune B, Grace JB, Urban DL (2002) MjM software design, Gleneden Beach, Oregon, USA.
- Mather PM (1976) Computational methods of multivariate analysis in physical geography. Wiley, London, UK.
- Kruskal JB (1964) Nonmetric multidimensional scaling: a numerical method. Psychometrika 29: 115-129.
- McCune B, Mefford MJ (2015) MjM Software, Gleneden Beach, Oregon, USA.
- Ramsey FL, Schafer DW (2002) The Statistical Sleuth. (2nd edtn), Duxbury, Pacific Grove, California, USA.
- St Clair JB, Mandel NL, Vance-Borland KW (2005) Genecology of Douglas Fir in Western Oregon and Washington. Ann Bot 96: 1199-1214.
- Wilhelmi N (2016) The Effects of Seed Source and Planting Environment on Douglas-fir (Pseudotsuga menziesii) Foliage Diseases. MS Thesis, Oregon State University, Corvallis, Oregon, USA.
- Wickham H (2016) ggplot2: Elegant Graphics for Data Analysis (Use R!). Springer, Dordrecht, Heidelberg, London, New York.
- Rajala T, Velmala SM, Vesala R, Smolander A, Pennanen T(2014) The community of needle endophytes reflects the current physiological state of Norway spruce. Fungal Biol118: 309-315.
 Spanish
Spanish  Chinese
Chinese  Russian
Russian  German
German  French
French  Japanese
Japanese  Portuguese
Portuguese  Hindi
Hindi