Research Article, Vegetos Vol: 30 Issue: 2
Present status of Fire Blight Caused by Erwinia amylovora on Quince Trees in the Republic of Macedonia
Julijana Arsova, Emilija Arsov and Sasa Mitrev*
Faculty of Agriculture, Goce Delcev University, Stip, Macedonia
*Corresponding author: Sasa Mitrev
Faculty of Agriculture, Goce Delcev University, Stip, Macedonia
Tel: 389/32-550-610
E-mail: sasa.mitrev@ugd.edu.mk
Received date: February 28, 2017 Accepted date: March 25, 2017 Published date: March 28, 2017
Citation: Arsova J, Arsov E, Mitrev S (2017) Present status of Fire Blight Caused by Erwinia amylovora on Quince Trees in the Republic of Macedonia. Vegetos 30:2. doi: 10.5958/2229-4473.2017.00139.2
Abstract
Erwinia amylovora, causal agent of fire blight disease, is an important and destructive bacterial pathogen, mainly on Rosaceae family such as pears, apples and quinces. Present status of E. amylovora, isolated from quince trees (Cydonia oblonga) from different localities in the Republic of Macedonia, included the period 2015-2016, are presented in this study. Severe occurrence of symptomatic quince threes, was recorded in almost all observing localities in Macedonia. After blooming period appeared first symptoms, most expressed symptoms were recorded during intensive growth of shoots. Some resulted from isolated strains elicted HR in tobacco, necrosis of pear seedlings and immature pear fruits. Strains have several biochemical characteristics typical and compare with positive control, such as: non-fluorescent, produce levan; oxidase negative, catalase positive; hydrolysis of gelatine positive; esculin and starch tests were negative; strains metabolise sucrose, sucrose, fructose and galactose, etc. Amplification of total DNA with PCR followed by agarose electrophoresis, originating from quince showed indistinguishable PCR patterns for all investigated native strains that were identical with European reference strains (LMG 1883). The aim of this study was to: i) check the status of E. amylovora on quince and ii) laboratory diagnostics and determination of isolated strains. The importance of the results is that disease incidence is sporadic and crop damages are of medium economic significance and it’s important to know that E. amylovora is present on quince only to be careful of serious outbreaks of fire blight on quince in Macedonia..
Keywords: Biochemical characteristics; Erwinia amylovora; Quince trees; PCR; Tobacco
Introduction
Fire blight, caused by Erwinia amylovora, is an old and most destructive disease of pears, apples and quinces and other pome fruit trees worldwide [1]. The disease causes significant and economic losses in many European and Mediterranean countries, but also in Macedonia the situation from the pass is very alarming and seriously [2]. In Macedonia, fire blight was also found in wild pear, for which in the past it means that genes for resistance for E. amylovora [3].
The bacteria can survive as an endophyte or epiphyte, and it’s survival time is dependent on various environmental factors [4]. E. amylovora can move quickly through host tissue, which can lead to the loss of an entire tree in a single growing season. The initial symptoms, which usually appear in early spring during warm and humid weather, started as a water-soaking of affected blossoms. These then wilted, shrivel and turned brown to black. Similarly, leaves wilt, shrivel, turn dark but remain attached to the tree for some time. Fruits can also be infected, appearing oily and often exuding droplets of bacterial ooze [5,6].
E. amylovora such a potentially destructive disease, is listed as an A2 quarantine organism by the European and Mediterranean Plant Protection Organization (EPPO) for Europe and the EPPO countries [3]. All the species are genetically homogeneous, and up till now no characteristics have been found that can clearly distinguish strains by geographical origin, original host or year of isolation [1,7]. This homogeneity has been confirmed with biochemical, serological and host range studies [1,3,8].
Material and Methods
Isolation and identification of the pathogen
In the period 2015/16, five different locality in the Republic of Macedonia were observed and symptomatic samples were collected for future analyses (Table 1). In the laboratory, suspect samples were surface sterilised with 0.5% sodium hypochlorite for 2.5 min and rinsed three times with sterile distilled water (SDW). Small pieces of these samples were placed in a sterile mortal and grounded with pestle in 0.5 mL of SDW.
| Nb | Isolate code | Year of isolation | Host plant | Region | Isolate |
|---|---|---|---|---|---|
| 1 | DvЕа-1 | 2015 | Cydonia oblonga | Valandovo, MK | Erwinia amylovora |
| 2 | DvЕа-2 | 2015 | Cydonia oblonga | Valandovo, MK | Erwinia amylovora |
| 3 | DvЕа-3 | 2015 | Cydonia oblonga | Valandovo, MK | Erwinia amylovora |
| 4 | DvЕа-4 | 2015 | Cydonia oblonga | Valandovo, MK | Erwinia amylovora |
| 5 | DvЕа-5 | 2015 | Cydonia oblonga | Valandovo, MK | Erwinia amylovora |
| 6 | DdEa-1 | 2015 | Cydonia oblonga | Delcevo, МК | Erwinia amylovora |
| 7 | DdEa-2 | 2015 | Cydonia oblonga | Delcevo, МК | Erwinia amylovora |
| 8 | DdEa -3 | 2015 | Cydonia oblonga | Delcevo, MK | Erwinia amylovora |
| 9 | DsЕа-1 | 2016 | Cydonia oblonga | Stip, MK | Erwinia amylovora |
| 10 | DsЕа-2 | 2016 | Cydonia oblonga | Stip, MK | Erwinia amylovora |
| 11 | DkЕа-1 | 2016 | Cydonia oblonga | Kocani, МК | Erwinia amylovora |
| 12 | DkЕа-2 | 2016 | Cydonia oblonga | Kocani,МК | Erwinia amylovora |
| 13 | DsrЕа-1 | 2016 | Cydonia oblonga | Strumica, МК | Erwinia amylovora |
| 14 | DsrЕа-2 | 2016 | Cydonia oblonga | Strumica, МК | Erwinia amylovora |
| 15 | DsrЕа-3 | 2016 | Cydonia oblonga | Strumica, МК | Erwinia amylovora |
| 16 | DsrЕа-4 | 2016 | Cydonia oblonga | Strumica, МК | Erwinia amylovora |
| 17 | Ks-550 (referent isolate) |
1991 | Pyrus communis | Kavadarci, MK | Erwinia amylovora |
| 18 | LMG1883 (referent isolate) |
1980 | Pyrus communis | Belgium | Erwinia amylovora |
Table 1: Referent isolate Macedonian samples compare with positive controls.
The resulting suspension was left to stand for 10 min, and then 100 μL was plated on sucrose nutrient agar (NAS) medium [9]. Inoculated plates were incubated at 25 ± 1°C until bacterial colonies developed on the plate surfaces (around 48 ours). E. amylovora like colonies were subcultured several times onto new NAS plates until pure cultures were obtained (white domed colonies). These isolates were kept at 4ºC for further identification. From total number of 56 isolates, only 16 were used for complete analyzes (Table 1). In addition, reference isolates were maintained in 20% glycerol at –80°C.
Growth of the bacterium
All strains were assayed for growth on sucrose nutrient agar (NAS) medium [9], King’s medium B agar (KB) [10] and on Crystal violet–cycloheximide–tergitol (CCT) medium [11]. NAS was used as a control for growth and levan production. The characteristics of the colonies grown on these media, were recorded 48 and 72 h after incubation and the experiment was repeated twice.
Pathogenicity test on immature pear fruit and leaves
Immature fruits and leaves of a highly susceptible cultivar, pear (Pyrus communis) 1.5–2 cm in diameter were collected, surface sterilised with 0.5% sodium hypochlorite and rinsed thrice with SDW. Each immature pear fruit and leaves, were stab inoculated with a needle using sterile destilate water as a negative control or a bacterial suspension at 109 colony forming units (CFU)/mL. This was prepared from a 24-h-old bacterial culture. Inoculated fruits and leaves were kept in a mist chamber at 26 ± 1°C and checked daily for the presence of necrosis accompanied by bacterial oozing, which was considered a positive indicator for E. amylovora. Leaves are stained in glass Erlenmeyer with sterile agar inside. The experiment was repeated twice for each isolate, using three replicates (three pear fruits and also leaves kept separately). The positive controls were E. amylovora Ks550 (Kavadarci, Mk) and LMG1883 (Belgium collection) (Table 1).
Hypersensitive response on tobacco
Bacterial isolates were grown overnight in liquid broth (LB) medium [12], pelleted cells were diluted in 1×PBS to 109 CFU/mL and infiltrated with a syringe into tobacco leaves (cv. Samsun). The plants were then incubated in a growth chamber at 25 ± 1°C with 16 h illumination and 80% humidity and readings were taken at 24, 48, 72 and 96 h. E. amylovora. LMG1883 and Ks550, were used as a positive controls and PBS was used as a negative control.
BIOLOG test
Positive bacterial isolates were grown and tested for the utilisation of 95 carbon sources, as available on the BIOLOG GN/GP MicroPlate system, using the previously mentioned BIOLOG method. The results from the BIOLOG system were analysed using the compatible software, Microlog™ version 4.20.05.
Chromosomal DNA isolation
For the molecular testing, a single colony from a 24-hour NA or NAS culture, which had been grown overnight at 25 ± 1°C, was suspended in 0.5 ml of molecular grade sterile water in a microcentrifuge tube, and directly used in the PCR mix [1].
PCR assays
The temperature of the initial denaturation step of the amplification cycle adequately lysed the bacterial cells, making the target genome accessible to the PCR primers and polymerases. The conventional (or single ) PCR was performed using primer sequences (primer A: 5’ CGG TTT TTA ACG CTG GG 3’ and primer B: 5’ GGG CAA ATA CTC GGA TT 3’), based on the plasmid pEA29 DNA and conditions described by Bereswill et al. [1]. The PCR reaction mixture (50 μl) contained a final concentration of 10 × PCR buffer, 10 mM of each dNTP, 50 mM MgCl2, formamide, 10 pmol μl-1 of each primer, 5 U μl-1 Taq polymerase and 5 μl of template DNA.
The reaction conditions were a denaturation step at 93°C for 5 minutes, followed by 40 cycles of 30 seconds each at 93°C for denaturation, then 30 seconds at 52°C for annealing, 72°C for 75 seconds for the extension step, and a final step for 10 minutes at 72°C to finish the reaction.
Results and Discussion
Isolation and identification of the causal agent
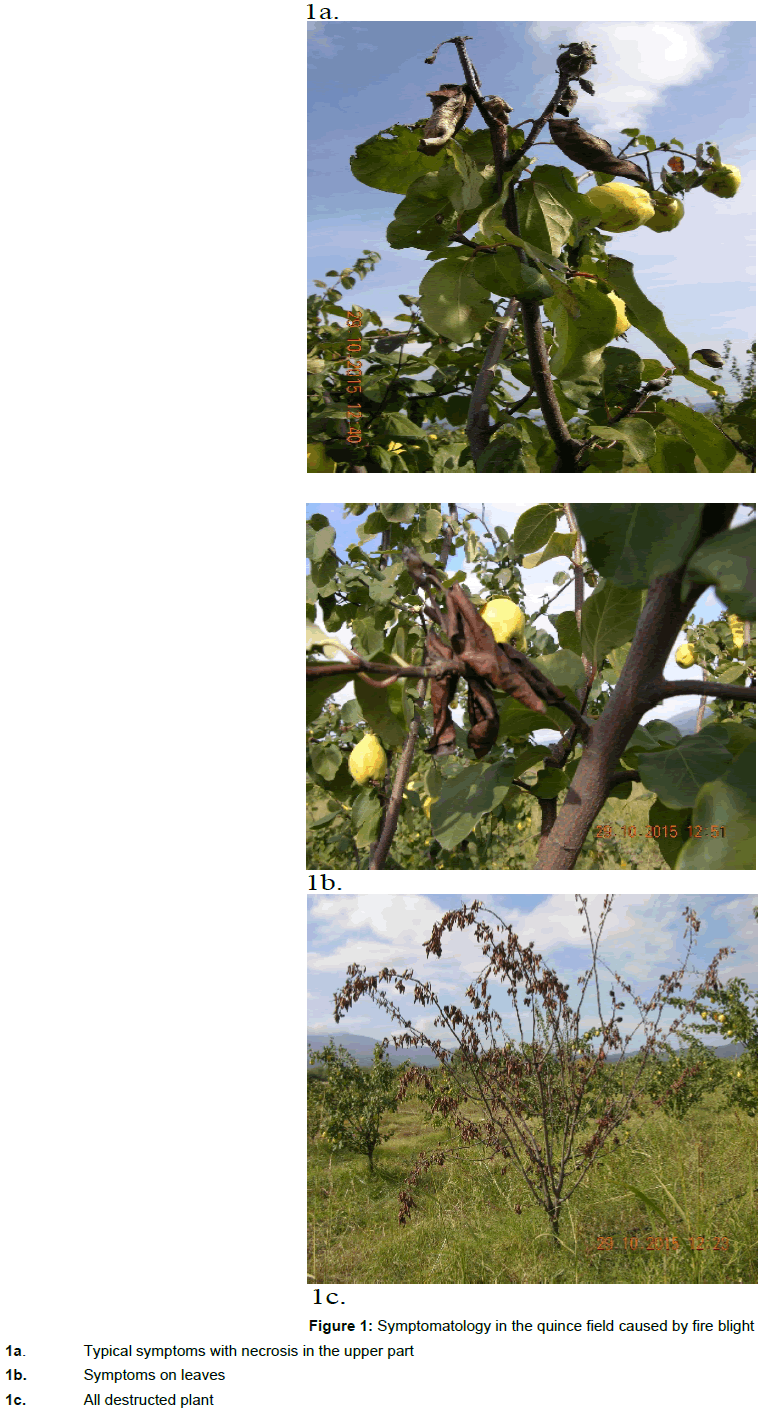
During the period 2015/16, several locality in the Republic of Macedonia were observed to check the situation with fire blight on quince (Cydonia oblonga). Symptomatology was typical for E. amylovora, in all parts and quince trees in Macedonia (Figure 1a, 1b, 1c). From total number of 56 isolates, 16 are laboratory tested for all analyzes, compare with positive control.
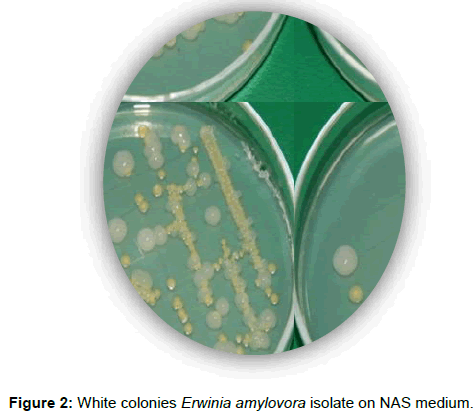
All samples first was isolate on NAS and they grow for 48h and show good levan type (Figure 2). After isolation, colony were kept on 4ºC and tested with biochemical and molecular analyzes.
Pathogenicity test on immature pear fruit and leaves
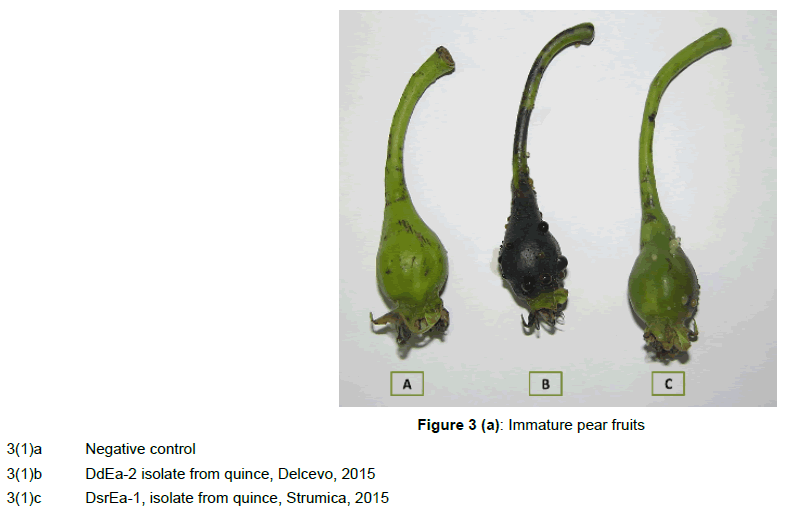
On immature pear fruits and leaves, the isolates induced necrosis accompanied with a yellowish bacterial exudate (oozing), which developed on the fruits after 3 days’ of incubation (Figure 3(1)a, 3(1) b, 3(1)c).
Hypersensitive response on tobacco
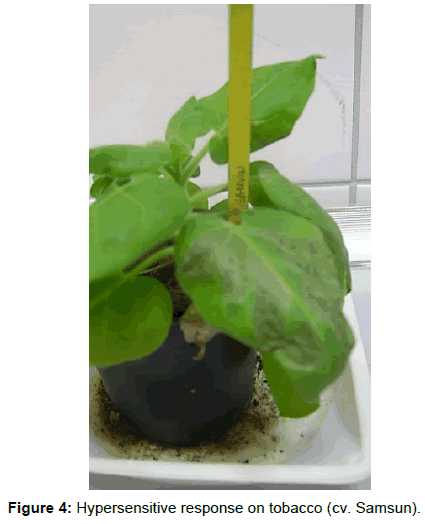
All 16 isolates tested developed a hypersensitive response (HR) on tobacco leaves (cv. Samsun). This occurred HR within 24 to 48 h and induced production of typical symptoms on leaves (necrosis) (Figure 4).
BIOLOG test
The results from the BIOLOG system were analysed using the compatible software, Microlog™ version 4.20.05, and showed 100% presence of E. amylovora (with similarities between 0.916 and 0.959) compared with the positive control (similarity of 0.993).
PCR amplification
The PCR products from the amplifications, a conventional (single) PCR, were separated on a 1.5% agarose gel in 0.5 × TAE buffer for 20 minutes at 120 V, then stained with 0.5 μg ml-1 ethidium bromide solution. The resulting gel was photographed under UV light.
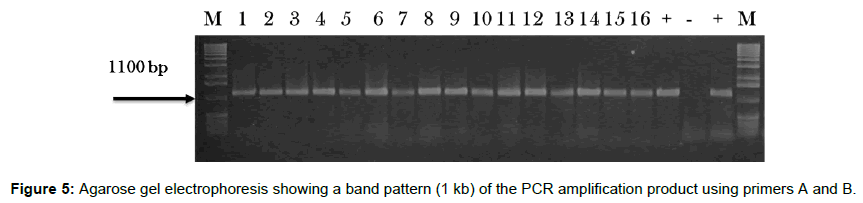
The region of the 16S rRNA gene (approximately 1100 bp) was amplified for each of the 16 Erwinia isolates using the universal primers A and B [13], compare with positive referente isolates (Ks550 & LMG 1883) (Figure 5).
The lane marked Ðœ contains the PCR markers ranging from 50- 2000 bp (Sigma); lanes 1– 16 contain isolates from quince shown in the Table 1; lane 17 contain positive control Ks550 isolate from Kavadarci, 1991; the lane marked B is a blank sample containing only molecular grade sterile water; the lane marked + contains the LMG 1883 representative positive control from Belgium; and the final lane marked M contains the PCR markers ranging from 50-2000 bp (Sigma).
Conclusions
Fire blight disease is an old and much observed diseases in apple, pear, quince trees and other plants from Rosacae family. The causal agent of the diseases Erwinia amylovora, is much known and laboratory tested. The aim of this study was to give an accent to the quince trees in the Republic of Macedonia, to check in the field what’s happening with bacterial status and what the possibility is for spreading the diseases.
The results from the symptoms in the field were recognised as typical for E. amylovora, and from total number of 56 isolates only 16 isolates were obtained for complete biochemical and molecular analyzes. The disease incidence is sporadic and crop damages are of medium economic significance and it’s important to know that E. amylovora is present on quince, to be careful of serious outbreaks of fire blight that can occur on quince in Macedonia. All isolates were shown to belong to E. amylovora based on morphology, culture, and biochemical, pathological and molecular characteristics, including PCR analyses of the 16S rRNA gene.
References
- Momol MT, Aldwinckle HS (2000) Fire blight in Moroccoand host range of Erwinia amylovora.. CABI 50: 55-72.
- Mitrev S (1995) Pathogenic and bacteriological characteristic Erwinia amylovora, pathogen on pear and quince trees in Macedonia. Plant Protection Journal 46: 97-109.
- Mitrev S, Kostadinovska E (2013) Wild pear - Pyrus pyraster as a new host of Erwinia amylovora in Macedonia. Journal of Plant Pathology 95: 167-170.
- Thomson SV (2000) Fire blight – the disease and its causative agent Erwinia amylovora’. CABI Publishing, USA.
- Vanneste JL, Eden-Green S (2000) Migration of Erwinia amylovorain host plant tissues. Fire blight – the disease and its causative agent Erwinia amylovora. CABI Publishing, Wallingford, USA.
- EPPO (2007a) EPPO A2 List of pests recommended for regulation as quarantine pests (version 2007-09).
- Vanneste JL (2000) What is fire blight? Who is Erwinia amylovora? How to control it? In ‘Fire blight – the disease and its causative agent Erwiniaamylovora’. CABI Publishing: Wallingford, USA.
- Vantomme R, Swings J, Goor M, Kersters K, Ley JDe (1982) Phytopathological, serological, biochemical and protein electrophoretic characterization of Erwiniaamylovorastrains isolated in belgium. Phytopathologische Zeitschrift103: 349-360.
- Billing E, Baker LA, Crosse JE, Garrett CM (1961) Characteristics of english isolates of erwinia amylovora(burrill). J Applied Bacteriol 24: 195-211.
- King EO, Ward MK, Raney DE (1954) Two simple media for the demonstration of pyocyanin and fluorescein. Journal of Laboratory and Clinical Medicine44: 301-307.
- Ishimaru C, Klos EJ (1984) New medium for detection of Erwinia amylovoraand its use in epidemiological studies.Phytopathology 74: 1342-1345.
- Miller TD, Schroth MD (1972) Monitoring the ephiphytic population of Erwinia amylovoraon pear with a selective medium. Phytopath 62: 1175-1182.
- Bereswill S, Pahl A, Bellemann P, Berger F, Zeller W, et al. (1992) Sensitive and species-specific detection of Erwinia amylovora by PCR-analysis. Appl Environ Microbiol 58: 3522-3526.
 Spanish
Spanish  Chinese
Chinese  Russian
Russian  German
German  French
French  Japanese
Japanese  Portuguese
Portuguese  Hindi
Hindi