Research Article, J Virol Antivir Res Vol: 14 Issue: 2
Estimation the Role of Interleukin-18 Promoter Polymorphisms (-607 C/A and 137 G/C) in Patient Infected with Hepatitis C Virus and Treatment with Sofosbuvir Drug
Mohey H Shikhoun1* and Omar AM Farghaly2
1Analyses and Laboratories Department, Higher Technological Institute of Applied Health Sciences in Sohag, Ministry of Higher Education, Cairo, Egypt
2Department of Biochemistry and Molecular Biology, Al-Azhar University, Asyut, Egypt
*Corresponding Author:Mohey H Shikhoun, Analyses and Laboratories Department, Higher Technological Institute of Applied Health Sciences in Sohag, Ministry of Higher Education, Cairo, Egypt; E-mail: moheyshikhoun@gmail.com
Received date: 17 October, 2024, Manuscript No. JVA-24-150264;
Editor assigned date: 21 October, 2025, PreQC No. JVA-24-150264 (PQ);
Reviewed date: 05 November, 2025, QC No. JVA-24-150264;
Revised date: 14 April, 2025, Manuscript No. JVA-24-150264 (R);
Published date: 21 April, 2025, DOI: 10.4172/2324-8955.1000720
Citation: Shikhoun MH, Farghaly OAM (2025) Estimation the Role of Interleukin-18 Promoter Polymorphisms (-607 C/A and 137 G/C) in Patient Infected with Hepatitis C Virus and Treatment with Sofosbuvir Drug. J Virol Antivir Res 14:2.
Abstract
Background: Two polymorphisms, -607 C/A and -137 G/C, together with their corresponding haplotypes, are known to affect the expression of Interleukin (IL-18). Numerous Single Nucleotide Polymorphisms (SNPs) in the gene promoter region influence IL-18 production. The study's objective is to ascertain whether IL-18 promoter polymorphisms (-607 C/A and 137 G/C) caused patients to respond or not during sofosbuvir treatment.
Methods: Using reverse transcriptase-PCR technology, patients in our study were split into two groups: Those who were anti-HCV positive and HCV-RNA negative (n=55) and responded to sofosbuvir treatment, and those who were anti- HCV positive and HCV-RNA positive (n=25) and did not respond to sofosbuvir treatment. Primer specific Polymerase Chain Reaction (PCR) for IL-18-137, 607 SNPs was performed for patients who responded and did not respond to sofosbuvir treatment.
Results: Our results show that of the 284 patients with hepatitis C virus infection, 258/284 (90.8%) responded to sofosbuvir treatment, while 26/284 (9.2%) did not. Out of the 284 patients who had the hepatitis C virus and were receiving sofosbuvir medication, we chose 80 individuals for additional testing, of whom 55 reacted to the medication and 25 did not. The IL-18 promoter SNPS-137 revealed that GG (56.4%) was higher in patients who responded to sofosbuvir therapy, while CC (68%) was higher in patients who did not respond to sofosbuvir therapy. In a similar vein, the IL-18 promoter SNPS-607 showed that patients who responded to sofosbuvir therapy had higher CC (63.6%) and those who did not had higher AC (52%).
Conclusion: This study shows how interleukin-18 polymorphisms affect how well Egyptian individuals respond to sofosbuvir treatment for chronic hepatitis C.
Keywords: Hepatitis C virus, Interleukin-18, Polymorphisms, Sofosbuvir
Introduction
Globally, the Hepatitis C Virus (HCV) affects roughly 185 million individuals [1]. Hepatocellular Carcinoma (HCC) and liver cirrhosis are two conditions that can result from HCV infection, which usually causes chronic liver damage [2]. HCV, a positive polarity enclosed single-stranded RNA virus, belongs to the genus Hepacivirus and family Flaviviridae [3,4]. The life cycle of HCV and host lipid metabolism are tightly connected, according to a growing body of research [5,6].
The pro-inflammatory cytokine IL-18 has a crucial role in controlling both the innate and acquired immune responses. Depending on the immunological milieu, IL-18 is implicated in Th1 and Th2 immune responses. IL-18 promotes the Th1-mediated immune response by inducing IFNG production when IL-12 is present; in the absence of IL-12, IL-18 promotes Th2 responses. In addition to being essential for the host's defense against intracellular microbial infection, IL-18 also triggers autoimmune disorders and prolongs the inflammatory process [7,8].
The IL 18 promoter polymorphism may affect how HCV infection progresses in some populations [9]. Three Single Nucleotide Polymorphisms (SNPs) at positions 137 G–C, -607 C–A, and -656 G– T are present in the IL-18 gene promoter. Because two of these, -607 C-A and -137 G-C, are located at the binding sites for the H4TFI nuclear factor and CREB transcriptional factors, respectively, cAMP response-element binding proteins and CREB transcriptional factors, mutations at these two sites may impact IL-18 expression and change the production of cytokines. Because IL-18 is an important immune regulator involved in the pathogenesis of HCV infection, we hypothesized that a promoter polymorphism known to modify IL18 expression and protein levels might influence the outcome of response to HCV-specific Sofosbuvir treatment. In this study, the role played by IL18 promoter polymorphisms _607C/A and _137G/C and their patterns of response and non-response to treatment with the drug sofosbuvir against hepatitis C virus were examined.
Materials and Methods
Ethics statement
Every patient involved in the study gave informed written consent; during regular clinic visits, blood samples were taken for various analyses, treatment plans and guidelines for enrollment in the national treatment program were announced by the Ministry of Health and approved by patients. Eighty individuals with hepatitis C virus were included in our study at random, without their interleukin-18 genotype being known. All samples were taken at Tahta Hospital in Egypt between November 2021 and June 2023 with consent from the hospital's governorate and patients.
Collection and processing of blood samples
A hepatitis C virus was tested on a sample of 1,715 people. Everyone lives in the Tahta center of the Sohag Governorate. During November 2021 and July 2023, each case was collected from Tahta Central Hospital. During that time, a brief investigation was carried out to identify hepatitis C viruses by using the ELISA method to detect anti-HCV antibodies. To identify IL-18 genotypes, a PCR-based method was improved. To collect blood samples, a cubital vein venipuncture was performed on each subject. The location was meticulously cleaned with 1% iodine and 70% isopropyl alcohol in water for one minute before being left to dry. To prevent the area from being contaminated, around 5 milliliters of blood were extracted using a sterile syringe and needle and then placed into clean plastic. The serum was extracted and stored at -70°C after the blood samples were centrifuged for ten minutes at 4000 rpm.
Our findings showed that out of 1715 people, 284/1715 (16.6%) had hepatitis C virus antibodies, while 1431/1715 (83.4%) had no antibodies at all. Our results revealed that of the 284 patients with HCV infection, 258/284 (90.8%) responded to sofosbuvir drug therapy, while only 26/284 (9.2%) did not.
Following the completion of the six-month treatment period, 55 patients who responded to antiviral therapy and 25 patients who did not were chosen. During routine clinic appointments, blood samples were collected for various analyses.
The patients in our study were divided into two groups: Those with anti-HCV positive and HCV-RNA negative patients (n=55) who responded to sofosbuvir therapy, and those with anti-HCV positive and HCV-RNA positive patients (n=25) who did not.
Serum markers for HCV infection
Anti-HCV antibody detection: Anti-HCV antibodies were detected using third-generation enzyme-linked immunosorbent assays (ELISAs; BioKit-bioelisa HCV 4.0). Recombinant antigens representing the major NS3, NS4, and NS5 epitopes of HCV were applied to the microplate wells. There is a correlation between the sample's HCV antibody levels and color intensity.
All of the chemicals were allowed to reach room temperature prior to starting the experiment. Liquid reagents were mixed prior to use. The concentrate washing solution was diluted one-tenth with distilled water. The concentrated conjugate was diluted 1/51 using the conjugate diluents. Diluted samples or controls were added to 96-well plates that had been pre-coated with a recombinant HCV-specific antigen. After that, the plate was incubated at 37°C for an hour to allow the Ag-Ab combination to form. The plate was cleaned, the conjugate was added, and it was then incubated for 30 minutes at 37°C. Tetra Methyl Benzidine (TMB) was introduced as a substrate solution for detection during the washing stage after the incubation time. The process was eventually stopped with H2SO4 , and the colorimetric signal was found by using a spectrophotometer to measure absorbance at 450 nm.
Molecular diagnostic
HCV RNA extraction: As directed by the manufacturer, viral RNA was extracted using the spin column technique and the viral RNA min kit. In summary, 560 µl of prepared AVL buffer containing carrier RNA and 140 µl of serum were pipetted into a 1.5 ml micro centrifuge tube, which was then incubated for 10 minutes at room temperature. Using a pulse-votexing approach, 560 µl of 97% ethanol was added to each sample and mixed for 15 seconds. The mixture was centrifuged for one minute at 8000 rpm after 635 µl of the pervious solution was carefully loaded onto the QIAamp spin column (in a 2 ml collection tube). A clean 2 ml collection tube with the QIAamp spin column was filled with 500 µl of AW1 buffer, and the tube was spun at 8000 rpm for one minute. The QIAamp spin column was repositioned in a cleaned 2 ml collection tube, 500 µL of buffers AW2 was added, and the tube was spun at full speed (14,000 rpm) for three minutes. Finally, 60 µl of AVE was added, centrifuged for one minute at 800 rpm, and allowed to equilibrate at room temperature for one minute. A complete HCV RNA sample was separated and collected in sterile vials for amplification.
Qualitative Polymerase Chain Reaction (PCR): Complementary DNA (cDNA) was produced for the first strand following the extraction of HCVRNA. Initial denaturation was done at 95°C for five minutes. Polymerase chain reaction amplification was carried out for one minute at 94°C, one minute at 57°C (annealing temperature), one minute at 72°C, and one minute at 72°C for the final extension for a total of forty cycles.
The primer sequences that were employed were 5′ACCCTCGTTTCCGTACAGAG 3′ for the reverse primer and 5′CGCGCGACTAGGAAGACTTC3′ for the forward primer.
Interleukin-18 genotyping
Prepration of Peripheral Blood Mononuclear Cells (PBMC): Five ml whole blood was withdrawn into an anticoagulant (EDTA) tube from each patient included in our study. The 5 ml blood sample was overlaid on the side wall of a centrifuge tube containing 2.5 ml of Ficoll-Hypaque density gradient separating solution under aseptic conditions. The blood on the Ficoll-Hypaque was centrifuged at 2000 rpm for 20 min at room temperature. After centrifugation the peripheral blood cellular components separated into 4 separate layers. The first layer was the plasma and platelets. The serum/plasma was aspirated carefully and kept frozen in a sterile vial at -80°C until tested for antibodies and other biochemical tests. The layer between the plasma and Ficoll-Hypaque at a density of 1.77 g/ml is the Peripheral Blood Mononuclear Cells (PBMC). The final layer was the erythrocytes. PBMCs were prepared from anti-coagulated blood from each subject in our study. Blood was diluted 1:1 with Dulbecco’s PBS, and PBMC prepared using Lymphocyte Separation Medium (LSM). After washing the harvested leucocyte-rich interphase in PBS, centrifuged and stored in a sterile vial at -80 áµ?C until amplification of IL-18 gene.
DNA extraction from PBMCs: The extraction kit, QIAamp DNA Mini, and blood for DNA purification from buffy coat spin column protocol all acquired from QIAGEN were utilized to extract total DNA from lymphocytes in order to identify IL-18. 20 μl of QIAGEN protease were pipetted into a 1.5 ml tube in order to extract DNA. Following that, we put 200 μl of the sample and 200 μl of buffer AL into the tube, and we incubated it for 10 minutes at 56°C. 200 microliters (96 percent) of ethanol were added to the tube and stirred for 15 seconds following a brief centrifuge. After adding the mixture to a 2 ml collection tube and centrifuging it for a minute at 8000 rpm, the QIAamp Mini spin column was used. A pristine 2 ml collection tube held the QIAamp Mini spin column. After that, we added 500 μl of buffer AW1 to the column and centrifuged it for a minute at 8,000 rpm. Next, we added 500 μl of buffer AW2 to the column and spun it for a minute at 14,000 rpm. After inserting the QIAamp Mini spin column into a fresh 1.5 ml micro centrifuge tube, we added 200 μl of buffer AE, allowed it to sit at room temperature for a minute, then spun it for another minute at 8,000 rpm.
Interleukin-18 genotypes determination: Wenham's previously published methods were used to determine the IL-18 genotypes. The QIAamp DNA micro kit was used to produce buffy coat DNA in accordance with the manufacturer's instructions. IL-18 genotypes were determined using a modified version of Wenham's PCR-based methodology. Sequence specific primer Polymerase Chain Reaction (PCR): Using PCR, the -137 SNPs were found. The forward primers 5'-CCCCAACTTTTACGGA AGAAAAG-3' and 5'- CCCCAACTTTTACGGAAG AAAAC-3' are common reverse primers, 5'-AGGAGGGCAAAATG CACTGG-3'. the 607 SNPs, the polymorphism was genotyped for IL-18 using RFLP analysis. Mutant primers were typically used in PCR genotyping, with the forward primer 5'GTTGCAGAAAGTGTAAAAAT TAGTA-3' and the reverse primer 5'-TAACCTCATTCAGGACT TCC-3'. Using deionized water, the PCR was carried out using 5 μl of genomic DNA isolated from lymphocytes, 25 pmol of each primer, and 25 μl of master mix. The total volume was then reached with 50 μl. A five-minute initial denaturation at 94°C was followed by forty cycles of denaturation at 94°C for thirty seconds, annealing at 65°C for thirty seconds, extension at 72°C for one and a half minutes, and final extension at 72°C for ten minutes in the thermal reactor. The 261-basepair (bp) amplified products underwent a 2-hour digestion process at 37°C.
Detection: Using a small prep electrophoresis chamber (Pharmacia, LKB) and 4% agrose gel electrophoresis with 10 μl of ethidium bromide (10 mg/ml), the PCR amplicon was found. The buffer containing tris-acetate EDTA was added to the electrophoresis tank. A 100 bp DNA marker (Gibco, BRL) was loaded along with a size detector, and 10 μl of the PCR result was carefully placed into the slot of the submerged gel after being combined with 10 μl loading buffer. Ethidium bromide fluorescence allowed for the visualization of the gel under UV light. The product sizes that were broken down by DraI for the _607 locations were 154 bp for the C allele and 125 and 28 bp for the A allele. The product sizes digested with BglII for the _137 locations were 141 bp for the C allele and 105 and 36 bp for the G allele.
Results
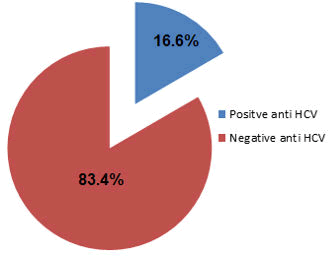
One thousand seven hundred and fifteen people were surveyed for hepatitis C virus by ELISA technique for anti-HCV antibody detection. Our findings indicated that 1715 people that there are 1715/284 (16.6%) patients with hepatitis C virus antibodies and 1715/1426 (83.4%) people completely devoid of hepatitis C antibodies by ELISA technique for anti-HCV antibody detection (Table 1 and Figure 1).
| No. Patients | Results | Percent % |
| 284 | Positive anti-HCV | 16.60% |
| 1426 | Negative anti-HCV | 83.40% |
Table 1: Detection of anti-HCV by ELIZA test on all collection samples.
Figure 1: Show the percentage of HCV infection in all collection samples.
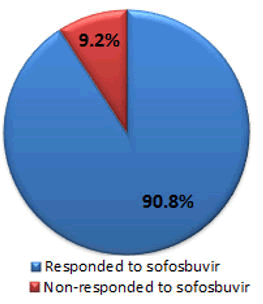
Our findings show that, of the 284 patients with HCV infection treated with sofosbuvir medication, 258/284 (90.8%) responded to treatment, while only 26/284 (9.2%) did not respond to treatment (Table 2 and Figure 2).
| No. Patients | Percent (%) | Responder to Sofosbuvir after treatment | HCV RNA by PCR |
| 258 | 90.80% | Responded | Negative |
| 26 | 9.20% | Non-responded | Positive |
Table 2: HCV RNA molecular detection for every patient following therapy.
Figure 2: Show the percentage of responded and non-responded after treatment to Sofosbuvir.
We selected 80 patients (55 patients who were respond to treated for the drug and 25 patients who did not respond to the treatment). Blood samples were taken during typical clinic visits in order to be analyzed in different ways. Two distinct patient groups were included in our study: Individuals whose patients are HCV-RNA negative and anti-HCV positive (n=55) who responded to sofosbuvir therapy, and those with anti-HCV positive and HCV-RNA positive patients (n=25) who did not respond to sofosbuvir therapy.
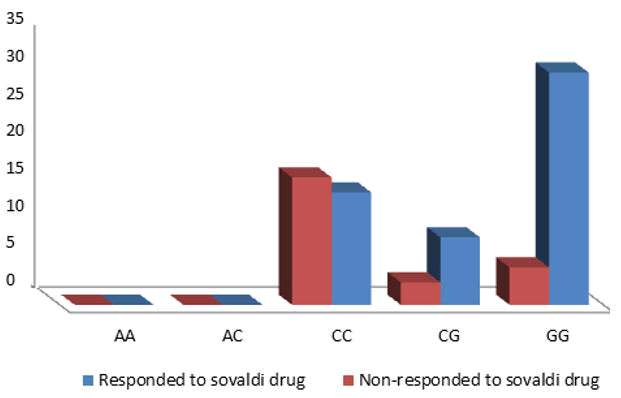
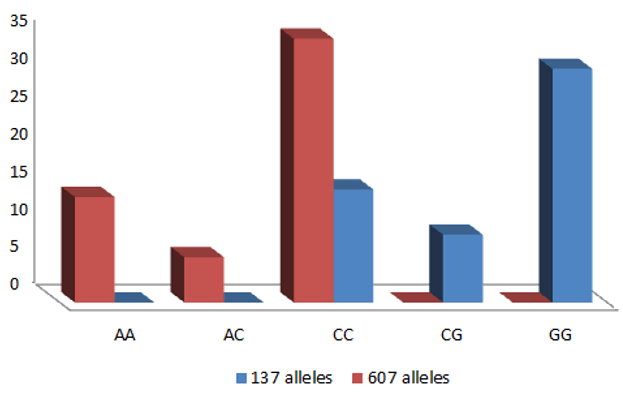
We discovered that the IL-18 promoter SNPS-137 shows high statistically significant difference between responded and non-responded patients to sofosbuvir therapy, IL-18 137 CC (27.3%), 137CG (16.4%), GG (56.4%) in responded patients to sofosbuvir therapy and 137 CC (68%), 137CG (12%), GG (20%) in non-responded patients to sofosbuvir therapy (Table 3 and Figure 3).
| Variables | Responded to Sofosbuvir drug (N=55) | Non-responded to Sofosbuvir drug (N=25) | X2 | P-value |
| AA | 0 | 0 | 12.396 | 0.002 H.S |
| AC | 0 | 0 | ||
| CC | 15 (27.3%) | 17 (68%) | ||
| CG | 9 (16.4%) | 3 (12%) | ||
| GG | 31 (56.4%) | 5 (20%) |
Table 3: Distribution of 137 alleles' IL-18 gene polymorphism for both response and non-response to sofosbuvir therapy.
Figure 3: Demonstrates the polymorphism variations' response and non-reaction toSofosbuvir therapy in relation to the 137 allele genotypes of the IL-18 gene promoter region.
Table 3 show that GG was higher among responded pateints to sofosbuvir therapy, while CC was higher among non-responded pateints to Sofosbuvir therapy with a Chi-square test result showing a statistically significant difference between.
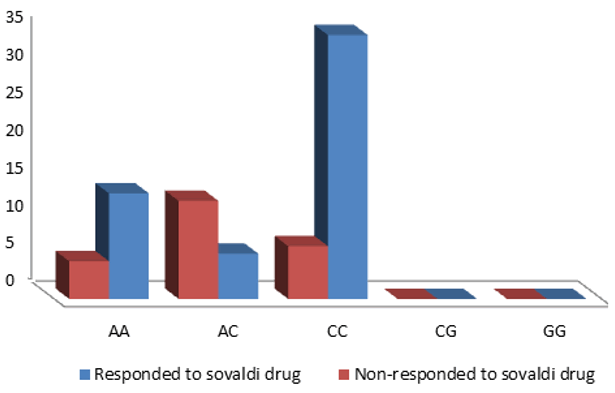
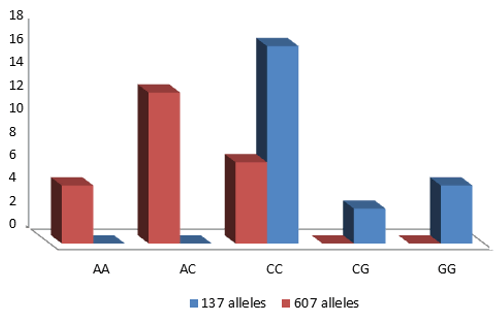
According to our findings, there is a highly statistically significant difference in the IL-18 promoter SNPS-607 between patients who responded and those who did not to sofosbuvir therapy, IL-18 607AA (25.5%), 607AC (10.9%), 607CC (63.6%) in responded patients to sofosbuvir therapy and 607AA (20%), 607AC (52%), 607CC (28%) in non-responded patients to Sofosbuvir therapy (Table 4 and Figure 4).
| Variables | Responded to sofosbuvir drug (N=55) | Non-responded to sofosbuvir drug (N=25) | X2 | P-value |
| AA | 14 (25.5%) | 5 (20%) | 16.592 | 0 H.S |
| AC | 6 (10.9%) | 13 (52%) | ||
| CC | 35 (63.6%) | 7 (28%) | ||
| CG | 0 | 0 | ||
| GG | 0 | 0 |
Table 4: Distribution of 607 alleles' IL-18 gene polymorphism for Sofosbuvir treatment response and non-response.
Figure 4: Demonstrates the polymorphism of the variations in response and non-response with solvaldi therapy as regarding promoter region of the IL-18 gene at genotypes 607 alleles.
Table 4 show that CC was higher among responded pateints to sofosbuvir therapy, while AC was higher among non-responded patients to sofosbuvir therapy with a Chi-square test result showing a statistically significant difference between.
According to our findings, the IL-18 polymorphism in response with solvaldi therapy shows high statistically significant difference between 137 alleles and 607 alleles, IL-18 137 CC (27.3%), 137CG (16.4%), 137GG (56.4%) and 607AA (25.5%), 607AC (10.9%), 607CC (63.6%) (Table 5 and Figure 5).
| 137 (N=55) | 607 alles (N=55) | X2 | P-value | |
| AA | 0 | 14 (25.5%) | 68 | 0 |
| AC | 0 | 6 (10.9%) | ||
| CC | 15 (27.3%) | 35 (63.6%) | ||
| CG | 9 (16.4%) | 0 | ||
| GG | 31 (56.4%) | 0 |
Table 5: Relationship between the patients' responded to Sofosbuvir therapy and IL-18 gene alleles 137 versus 607.
Figure 5: Shows the polymorphism variations among patients' responses to sofosbuvir therapy with respect to the IL-18 gene promoter region at genotypes 607 and 137 alleles.
Table 5 demonstrates a highly statistically significant difference between both alleles in patients who responded to sofosbuvir therapy by using Chi-square test.
According to our findings, the IL-18 polymorphism in patient’s nonresponse with solvaldi therapy shows high statistically significant difference between 137 alleles and 607 alleles, IL-18 137CC (68%), 137CG (12%), 137GG (20%) and 607AA (20%), 607AC (52%), 607CC (28%) (Table 6 and Figures 6-8).
|
|
137 alles (N=25) |
607 allles (N=25) |
X2 |
P-value |
|
AA |
0 |
5 (20%) |
30.167 |
0 |
|
AC |
0 |
13 (52%) |
||
|
CC |
17 (68%) |
7 (28%) |
||
|
CG |
3 (12%) |
0 |
||
|
GG |
5 (20%) |
0 |
Table 6: Relationship between the patients' non-responded to Sofosbuvir therapy and IL-18 gene alleles 137 versus 607.
Table 6 demonstrates a highly statistically significant difference between both alleles in patients who non-responded to sofosbuvir therapy by using Chi-square test.
Figure 6: Shows the polymorphism variations among patients' nonresponses to sofosbuvir therapy with respect to the IL-18 gene promoter region at genotypes 607 and 137 alleles
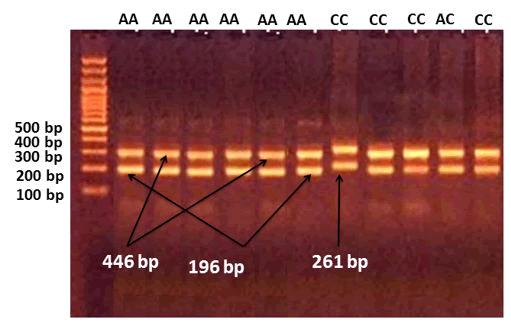
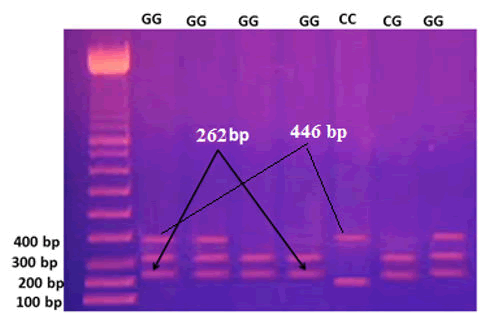
Figure 7: An illustration of the Interleukin-18 (IL-18)-137 sequence-specific amplification–polymerase chain reaction product analysis obtained using agarose gel electrophoresis. In contrast, for the M marker (100–1,200 bp), the presence of the G or C allele was found at a 261-bp product size. The (GG) homozygote of the wild type was only visible in the G allele, the (CC) homozygote of the mutant type was only visible in the C allele, and the (G/C) heterozygote was detectable in both the G and C alleles. internal control for a product size of 446 bp.
Figure 8. The agarose gel electrophoresis picture shows the results of sequence-specific amplification-polymerase chain reaction product analysis of Interleukin-18 (IL-18) –607. In contrast, at 196 bp product size where M is the marker presence of the C or A allele was found (100–1,200 bp). The (CC) wild type homozygote was only visible in the C allele, the (AA) mutant type homozygote was only visible in the A allele, and the (C/A) heterozygote was visible in both the C and A allele, with internal control at 446 bp product size.
Discussion
As a proinflammatory cytokine, IL-18 causes liver cells to become inflamed directly. TNF- α and IL-18 have been shown to be crucial mediators of T cell-mediated liver damage in animal tests. Simultaneous neutralization of TNF-α and IL-18 completely shielded mice from exotoxin-induced hepatic injury. The liver damage caused by LPS was entirely avoided by using an anti-IL-18 neutralizing antibody. In clinical investigations, it was discovered that IL-18 was connected to hepatic damage and was up-regulated during chronic hepatitis C.
Single Nucleotide Polymorphisms (SNPs) in the gene may impact the synthesis of Interleukin-18 (IL-18), which is necessary for the clearance of viruses and the development of disease.
In the current investigation, we found that IL-18 promoter SNPS-137 show high statistically significant difference between responded and non-responded patients to Sofosbuvir therapy, where GG was higher among responded patients to sofosbuvir therapy, and CC was higher among non-responded patients to sofosbuvir therapy.
On the contrary in this study, we found that Il-18 promoter SNPS-607 show high statistically significant difference between responded and non-responded patients to Sofosbuvir therapy, where CC was higher among responded patient's to Sofosbuvir therapy, while AC was higher among non-responded patient's to Sofosbuvir therapy.
According to Khripko et al., the group with lower levels of enhanced IL-18 production has a higher frequency of the C allele. this finding is in agreement with our results that the IL-18-607 CC genotyping was 63.0% in responded patients to sofosbuvir, while IL-18 137 CC genotyping was 68.0% in non-responded patients to Sofosbuvir.
Our findings are agreement with the hypothesis put forth by Giedraitis et al. that two transcription binding sites, a transversion G to C at position -137 and a transversion C to A at position -607, are modified by single-nucleotide polymorphisms of IL-18 and affect the quality of transcribed IL-18 mRNA.
Our findings demonstrated a strong statistically significant difference between 137 and 607 alleles in the IL-18 polymorphism in patients who do not respond to Solvadi therapy: 137 CC (68%), 137CG (12%), 137GG (20%) and 607 AA (20%), 607AC (52%), 607CC (28%).
Conclusion
IL-18 polymorphism is linked to the ability of mononuclear cells to generate IL-18. There existed two polymorphisms in high linkage disequilibrium: 137 G/C and 607 C/A. Our findings demonstrated a strong statistically significant difference between 137 and 607 alleles in the IL-18 polymorphism in patients who do not respond to Solvadi therapy: 137 CC (68%), 137CG (12%), 137GG (20%) and 607 AA (20%), 607AC (52%), 607CC (28%).
Funding
There was no funding for this study.
Data Availability
The datasets used and analyzed during the current study are available from the corresponding author on reasonable request.
Declarations
Approval was obtained from the Ethics Board of the Faculty of Pharmacy, Al-Azhar University for the study. All patients signed an informed consent before participating in the study.
Consent for Publication
Not applicable.
Competing Interests
The authors declare no competing interest.
References
- Mohd Hanafiah K, Groeger J, Flaxman AD, Wiersma ST (2013) Global epidemiology of hepatitis C virus infection: new estimates of age-specific antibody to HCV seroprevalence. Hepatology 57: 1333-1342.
[Crossref] [Google Scholar] [PubMed]
- Barth H, Liang TJ, Baumert TF (2006) Hepatitis C virus entry: molecular biology and clinical implications. Hepatology 44: 527-535.
[Crossref] [Google Scholar] [PubMed]
- Huang H, Sun F, Owen DM, Li W, Chen Y, et al. (2007) Hepatitis C virus production by human hepatocytes dependent on assembly and secretion of very low-density lipoproteins. Proc Natl Acad Sci USA 104: 5848-5853.
[Crossref] [Google Scholar] [PubMed]
- Zeisel MB, Fofana I, Fafi-Kremer S, Baumert TF (2011) Hepatitis C virus entry into hepatocytes: molecular mechanisms and targets for antiviral therapies. J Hepatol 54: 566-576.
[Crossref] [Google Scholar] [PubMed]
- Kim DS, Burt AA, Ranchalis JE, Jarvik ER, Rosenthal EA, et al. (2013) Novel gene-by-environment interactions: APOB and NPC1L1 variants affect the relationship between dietary and total plasma cholesterol. J Lipid Res 54: 1512-1520.
[Crossref] [Google Scholar] [PubMed]
- Nakanishi K, Yoshimoto T, Tsutsui H, Okamura H (2001) Interleukin-18 regulates both Th1 and Th2 responses. Annu Rev Immunol 19: 423-474.
[Crossref] [Google Scholar] [PubMed]
- Gracie JA, Robertson SE, McInnes IB (2003) Interleukin-18. J Leukoc Biol 73: 213-224.
[Crossref] [Google Scholar] [PubMed]
- An P, Thio CL, Kirk GD, Donfield S, Goedert JJ, et al. (2009) Regulatory polymorphisms in the interleukin-18 promoter are associated with hepatitis C virus clearance. J Infect Dis 198: 1159-1165.
[Crossref] [Google Scholar] [PubMed]
- Giedraitis V, He B, Huang WX, and Hillert J (2001) Cloning and mutation analysis of human IL-18 promoter: a possible role of polymorphisms inexpression regulation. J Neuroimmunol 112: 146-152.
[Crossref] [Google Scholar] [PubMed]
 Spanish
Spanish  Chinese
Chinese  Russian
Russian  German
German  French
French  Japanese
Japanese  Portuguese
Portuguese  Hindi
Hindi