Research Article, J Virol Antivir Res Vol: 6 Issue: 3
Examination of Infection Parameters for Replication of Israeli Isolate of Koi Herpesvirus in Common Carp Brain Cells
Lisa Katharina Jordan1, Anette Amtmann1, Elke Heidenreich1, Jürgen Christian2, Rainer Buchholz1 and Anna Maria Becker1*
1Institute of Bioprocess Engineering, Friedrich-Alexander-University Erlangen- Nürnberg, Paul-Gordan-Straße 3, 91052 Erlangen, Germany
2Bavarian Health and Food Safety Authority, Institute for Animal Health II, Eggenreuther Weg 43, 91058 Erlangen, Germany
*Corresponding Author : Anna Maria Becker
Institute of Bioprocess Engineering, Friedrich-Alexander-University Erlangen-Nürnberg, Paul-Gordan-Straße 3, 91052 Erlangen, Germany
Tel: +49 9131 8523054
Fax: +49 9131 85 23057
E-mail: anna.maria.becker@fau.de
Received: October 14, 2017 Accepted: October 23, 2017 Published: October 30, 2017
Citation: Jordan LK, Amtmann A, Heidenreich E, Christian J, Buchholz R, et al. (2017) Gene Cloning, Examination of Infection Parameters for Replication of Israeli Isolate of Koi Herpesvirus in Common Carp Brain Cells. J Virol Antivir Res 6:3. doi: 10.4172/2324-8955.1000176
Abstract
Research on koi herpesvirus (KHV) is mainly focused on fish trials for the investigation of virus entry, disease progress and immunological responses within the organism in order to develop new diagnostic and treatment strategies. However, examination of in vitro-replication of KHV might give useful insights supporting the understanding of this pathogen as well as contributing to the development of new methods for diagnostic purposes and/or vaccine development. Nevertheless, until today, only few studies have been published on this matter.
Thus, this work concerns investigation of in vitro-replication of the Israeli isolate of KHV, KHV-I, in the widely used common carp brain cells (CCB) to identify the parameters leading to high-quality virus stock with high titers for further applications. Therefore, the propagation of the KHV-I at the influence of various cell densities at inoculation, virus load and time of harvest on the propagation of KHV-I was examined.
Obtained results indicate that inoculation with relatively high cell densities is advantageous since the highest titers could be reached for 60 000 and 75 000 cells/cm2 already 3 days after inoculation. In contrast, the use of too high virus load for inoculation with too low cell density can result in much lower maximum or even decrease in virus titers. While viral DNA showed to be relatively stable after reaching the maximum, the infectivity of the virus can decrease shortly after the peak. Therefore, selection of the right combination of infection parameters is crucial for reaching the appointed goal.
This work gives valuable insights into the replication of KHV-I in CCB cells and indicates the differences between various KHV isolates regarding in vitro-propagation.
Keywords: Common carp brain cells; Koi herpesvirus; Multiplicity of infection; Time of infection; Time of harvest; Replication; Virus titer
Introduction
Koi herpesvirus (KHV), also known as Cyprinid herpesvirus-3 (CyHV-3), is a DNA virus belonging to the family Alloherpesviridae in the order Herpesvirales [1]. The virus that causes the koi herpesvirus disease (KHVD) in carp was first reported in Israel after outbreaks in several farms [2]. Meanwhile, this viral pathogen is widely spread and an issue in several parts of the world, e.g. in the USA, Poland, China and Japan [3-8]. As all known herpes viruses, it can undergo a latency phase in the infected host until environmental parameters change and trigger replication [9,10]. Sick fish exhibit characteristic symptoms, e.g. skin lesions, bilateral enophtalmos and necrotic gill tissue [5,11,12]. The virus is believed to enter the organism via skin [13] or gills [11,14,15].
It is suggested that, on cellular level the virus enters cells via lipid rafts in the cell membrane and that cholesterol plays an important role [16]. Based on in vitro-replication experiments and electron microscopy, abnormal virus particles which do not exhibit a core or show chromatin attached to the particle membrane could be observed for KHV. However, if these particles are functional and can confer infection is questionable [6,17,18]. On the other hand, also nonenveloped viral particles can be produced during the replication, e.g. due to the lysis of the cell prior to the release of the virus particles or due to prevented budding by unfavorable modification of membrane proteins, as already reported, e.g. for hepatitis B and hepatitis C virus [19-21]. As the envelope is crucial for the virus attachment and penetration of the cell membrane [19] such incomplete virions also contribute to demeaned replication of the virus. Thus, the efficiency of the in vitro-replication of a given virus stock depends on the number of infective virus particles in relation to such that cannot cause infection [22,23].
There are a number of methods that can be used for both diagnostic and research purposes in work with the KHV. Whereas real-time PCR, e.g. according to Gilad et al. [15] or Bergmann et al. [4], can be used for very sensitive detection of viral DNA, in vitro replication in combination with 50 % tissue culture infective dose (TCID50) and statistical evaluation allows estimation of the titer of infective viral particles [24,25]. Combining the data of these both techniques can lead to new insights and valuable information about in vitro-replication of this pathogen at various conditions.
Nevertheless, so far, only few groups were able to report titers of this virus that extend 104 TCID50 or plaque forming units (PFU) per mL of cell culture medium after in vitro-replication [26-29], suggesting that either the cell lines that are currently available for inoculation of KHV are not able to propagate the virus to higher titers and/or parameters of the infection and propagation process are not optimal. Furthermore, different isolates of KHV seem to vary regarding their replication: whereas the Taiwanese isolate, KHV-T, can reach titers as high as 107 TCID50/mL [25,29], lower titers of 103 to 106 PFU/mL were reported by Ronen et al. [26] for KHV-I. The analysis of DNA from various isolates showed that most differences are due to frameshift mutations that can lead to varying glycoproteins exhibited in the envelope [30]. As the envelope is crucial for virus entry to the cell, this might be a reason for less virulent strains and/or their slower replication. However, further research is needed in order to reach a conclusion.
Mletzko et al. [25] showed that adjusting infection parameters such as: time of infection (TOI, expressed as cell density at inoculation), multiplicity of infection (MOI, number of infective virus particles per cell used for inoculation) as well as time of harvest (TOH) can lead to reproducibly high titers for KHV-T in the widely used common carp brain cells (CCBs), reaching 108 TCID50/mL. Moreover, only few publications are available in which comparably high titers were reported [29]. However, above mentioned infection parameters are often neither investigated nor adjusted for efficient replication.
Thus, the aim of this study was to investigate the influence of TOI, MOI and TOH on in vitro-replication of KHV-I, that is one of the most prevalent isolate of this virus, in order to reach the best possible replication. As CCB cells are widely used for KHV replication and high virus titers could previously be reached for other isolates, this cell line was also used in this study. An efficient and robust replication process is necessary to obtain enough virus particles for further examination of this isolate without laborious pre-concentration and purification. Moreover, characterization of in vitro-replication can lead to new advances for disinfection, diagnostics and vaccine production for KHV.
Materials and Methods
Cells and virus
The cell line from common carp brain used in this work was received in passage 83 from Friedrich-Loeffler Institute (CCB, Cat. No. CCLV-RIE 816, Lot. No. 4223, FLI Insel Riems, Greifswald, Germany). Strain maintenance was performed using tissue flasks for adherent cells (T-75 flasks, Sarstedt, Kleinstadt, Germany) and Eagle’s Minimum Essential Medium with Earle’s salts (EMEM; Sigma, Munich, Germany), supplemented with 1x non-essential amino acids (Biochrom, Berlin, Germany), 9 % fetal bovine serum (FBS, Biochrom, Berlin, Germany) and 25 mM HEPES (Carl Roth, Karlsruhe, Germany). An addition of 26 mM NaHCO3 (Carl Roth, Karlsruhe, Germany) to the medium, provided neutral pH in an atmosphere of 5 % CO2. The virus, CyHV-3-I (KHV-I, Reg. No. RVB- 403), was received from Prof. Dr. Jens Peter Teifke (FLI, Insel Riems, Greifswald, Germany) in passage 3 and was originally isolated from adult koi from Israel [5]. Virus stock used for further experiments was prepared with CCB cells, seeded in EMEM, supplemented with 9 % FBS, 25 mM HEPES and 0.5 mM NaHCO3 in three T-75 flasks with a cell density of 60 000 cells/cm2. The day after, the medium was discarded and the cells were inoculated with 3 mL of a diluted KHV-I virus stock achieved prior, and incubated at 25°C for one hour. Next, high glucose Dulbecco’s Minimum Essential Medium (DMEM, Life Technologies, Carlsbad, USA), supplemented with 9 % FBS, was added to a total volume of 20 mL, the flasks were incubated at 25°C and transferred to – 80°C for harvest 5 days post infection (d p.i.). The virus suspension of these flasks was pooled after thawing, aliquoted (1-5 mL) and the so obtained stocks transferred to – 80°C. The virus titer of the virus suspension was analyzed via TCID50 and qPCR and used for following infection experiments.
Experiments
Experiments with various cell densities at virus inoculation and virus titer were performed in order to appoint the infection parameters leading to the maximum virus titers. Moreover, various time points for harvest were chosen to illustrate infection course at investigated parameters. For this purpose, different TOI (cell densities at inoculation in cells/cm2) of 15 000, 30 000, 60 000 and 75 000 cells/ cm2 were seeded in T25-flasks in EMEM and incubated at 25°C with an atmosphere of 5 % CO2. In order to assure reproducible and reliable adjustment of TOI, cell counts of detached cells were performed prior seeding in counter chambers (LO – Laboroptik, Lancing, UK) using trypan blue assay (Sigma, Steinheim, Germany). After 1 d of incubation, the medium was discarded and the attached cells were incubated with 1 mL virus suspension with adjusted virus titer for 60-80 minutes. Afterwards, fresh DMEM was added to a total volume of 7 mL and the flasks were incubated at 25°C. To estimate the exact MOI with that the cells were infected, an aliquot of the used virus suspension was analyzed via TCID50. The investigated MOI were 0.0002, 0.0007, 0.003, 0.006 and 0.012. Furthermore, a modified qPCR, based on Gilad et al. [15], was performed to determine the viral DNA load in harvested virus suspensions. Samples were examined microscopically prior to the harvest, that was performed at day 3, 4, 5, 6, 7, and 10 p.i., or, 1, 2 and 3 d p.i., by placing culture flasks at – 20°C for at least one day. Each time point corresponds to one culture flask.
TOI lower than 15 000 cells/cm2 were not tested since seeding CCB cells at such densities proved previously to be impracticable, due to absent cell-cell-connection or insufficient amount of growth factors present. Additionally, higher TOI than 75 000 cells/cm2 were excluded from the experiments because of incapability of the cells to attach when seeded at higher densities [25].
Monitoring of virus replication
Virus titers were estimated via TCID50, as described previously by Mletzko et al. [25], with slight modifications: CCB cells were seeded in 96-well plates (Greiner Bio-One, Frickenhausen, Germany) with a cell density of 40 000 cells/cm2 in DMEM (as described above) and incubated for one day at 25°C. Next, the harvested virus samples were thawed and diluted in 10-fold dilutions up to 8 dilution steps, whereas each dilution was applied to 8 wells. The plate was sealed, incubated at 25°C for 12 days and finally evaluated for signs of virus infection. Presence of syncytia and/or cytopathic effect was considered as positive signal of virus infection.
For analysis of viral DNA load, samples were introduced to qPCR (CFX Connect Real Time System, Bio-Rad, Hercules, USA.) analysis as described by Gilad et al. [15] after extraction with the Qiagen DNeasy Blood and Tissue Kit (Qiagen, Venlo, the Netherlands).
Mathematical processing of the data
In order to simplify discussion of the results, the efficiency of replication was evaluated by forming the ratio of the virus titer at maximum virus infectivity during the infection course to the virus titer used for inoculation according to the formula:
Furthermore, to discuss the maximum virus yield per cell, following calculation was used:
where Vm is the total volume of culture medium during replication and F is the surface of the used culture flask, e.g. 25 cm2 for T-25.
Results
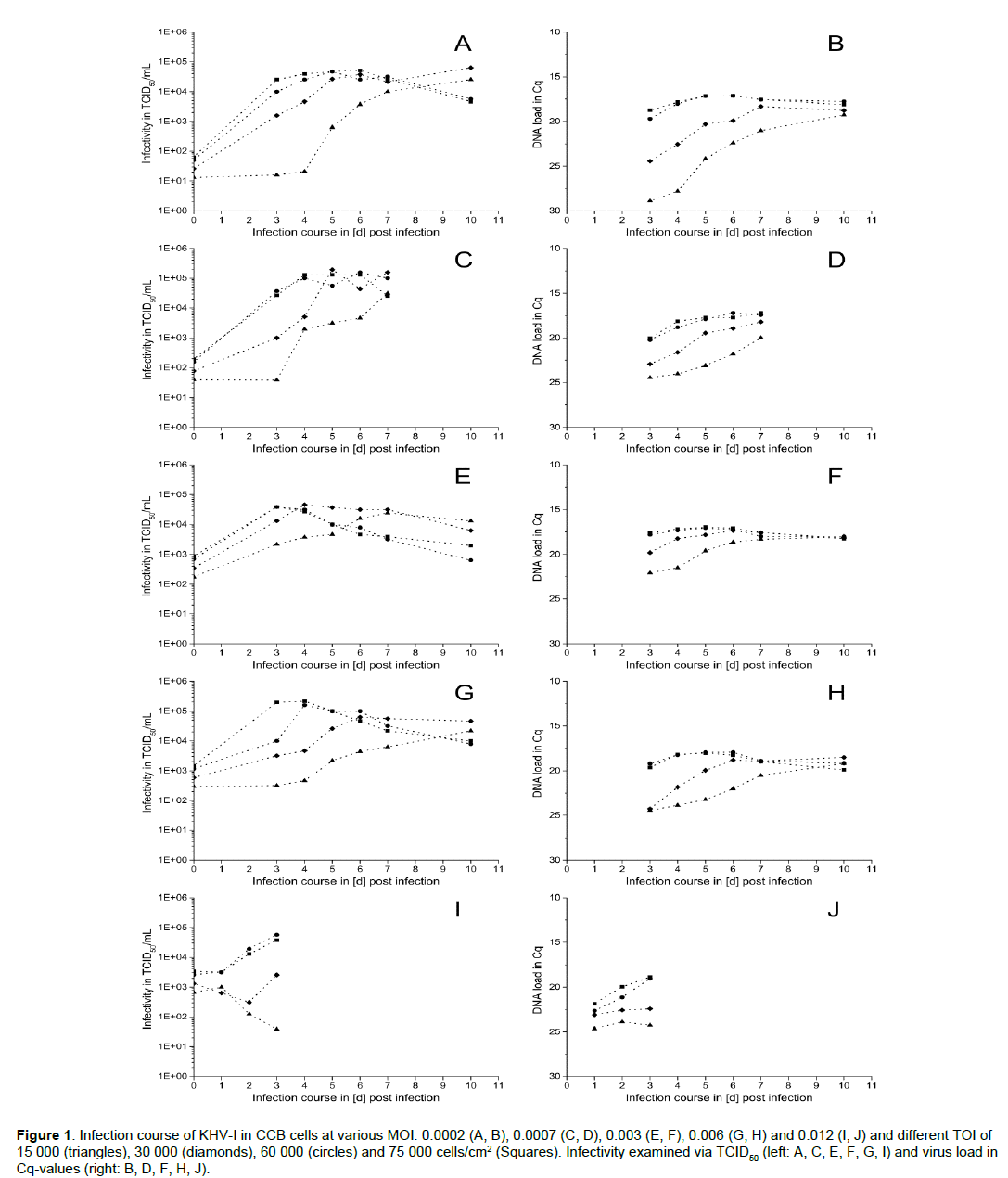
The infection progress of KHV-I in CCB cells with various TOI and MOI, expressed as virus titer by TCID50/mL (left) and viral DNA load (right) at various time points is shown in Figure 1. Independent of the applied MOI, the highest titers (up to 5 × 104 TCID50/mL) were reached for TOI between 30 000 and 75 000 cells/cm2, whereas the virus titer for the lowest cell density (15 000 cells/cm2) reached comparable values only at the end of the experiments (10 d p.i.) (Figure 1A). Furthermore, using higher TOI resulted in faster replication of the virus, thus causing a shift of virus titer peak. Where for TOI of 30 000 cells/cm2, except for MOI 0.003, the highest virus titers were reached between day 5 and 7 p.i., for 60 000 and 75 000 cells/cm2 the peaks were registered already at day 4 p.i. Furthermore, while for the highest cell densities virus titer started to decrease from day 6 p.i. on, the viral DNA-load (expressed as Cq-values) did not show similar reduction (Figure 1, right). In contrast, for TOI of 60 000 and 75 000 cells/cm2 viral DNA-load showed only slight fluctuations between day 3 and 10 p.i. after reaching the maximum. For the two lower TOI that were examined here, an increase between day 3 and 7 p.i. in viral DNA load was observed, correlating well with the increasing TCID50-values.
Figure 1: Infection course of KHV-I in CCB cells at various MOI: 0.0002 (A, B), 0.0007 (C, D), 0.003 (E, F), 0.006 (G, H) and 0.012 (I, J) and different TOI of 15 000 (triangles), 30 000 (diamonds), 60 000 (circles) and 75 000 cells/cm2 (Squares). Infectivity examined via TCID50 (left: A, C, E, F, G, I) and virus load in Cq-values (right: B, D, F, H, J).
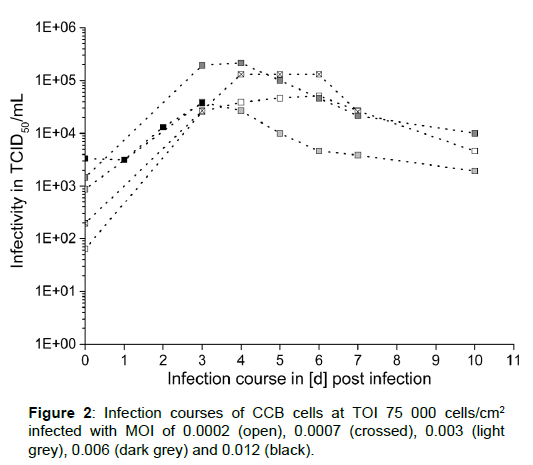
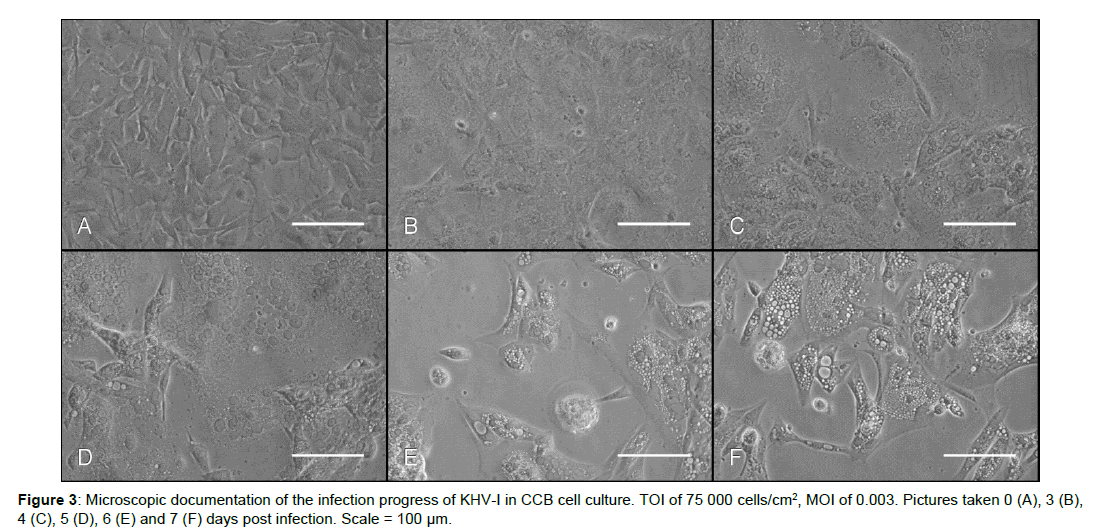
Comparing virus titers reached at constant TOI inoculated with varying MOI, it can be noticed that nearly the same maximum concentrations of infective virus particles were registered for all tested MOI values (Figure 1). Using the lowest TOI (15 000 cells/ cm2) and the highest MOI (0.012) a decrease instead of an increase of the titer was observed between day 1 and 5 (Figure 1I). A combination of 30 000 cells/cm2 with the same MOI resulted in a lag-phase up to day 2 p.i. followed by only slight increase at day 3 p.i.. In contrast, viral DNA-load stayed more or less constant in these infection experiments (Figure 1J, right). At higher TOI of 60 000 and 75 000 cells/cm2, the highest virus titers of around 2 × 105 TCID50/mL were reached for MOI of 0.0007 and 0.006. For better comparison the data for TOI of 75 000 cells/cm2 and all investigated MOI values is shown in Figure 2. With an exception of the highest investigated MOI (0.012), the virus titer reached its maximum between day 1 and 3 either followed by slight fluctuation in TCID50 values (at MOI of 0.0002 and 0.0007) or even decrease (at MOI of 0.003 and 0.006). At the same time, corresponding DNAload showed only slight fluctuations. Additionally, microscopic documentation of the infection progress for TOI of 75 000 cells/cm2, inoculated with MOI of 0.003 is shown in Figure 3. Whereas symptoms of infection, such as cells with prominently enlarged nuclei, could already be observed 3 d p.i. throughout the whole cell layer, only little plaques and sporadic syncytia forming were visible. Interestingly, the maximum virus titer occurred already at this point and started to decrease one day later. As replication of the virus progressed, visual symptoms of infection became more obvious: infected cells developed intracellular vacuoles, severe plaque formation was observed 6 and 7 d p.i., followed by cell detachment and lysis.
When looking at the virus replication efficiency, it can be noticed that the highest virus yield was obtained for the lowest MOI values independent of the TOI (Table 1), even if the maximum virus titer was not as high as when using higher MOI.
| TOI [cells/cm2] | MOI 0.0002 | MOI 0.0007 | MOI 0.003 | MOI 0.006 | MOI 0.012 |
|---|---|---|---|---|---|
| 15 000 | 3.3 | 2.9 | 2.1 | 1.9 | 0.2 |
| 30 000 | 3.4 | 3.4 | 2.1 | 2.0 | 0.3 |
| 60 000 | 3.0 | 3.0 | 1.7 | 2.1 | 1.3 |
| 75 000 | 2.9 | 2.8 | 1.6 | 2.2 | 1.0 |
Table 1: Virus replication efficiency, expressed as ratio of the virus titer at maximum virus infectivity during the infection course to the virus titer used for inoculation (according to Equation 1).
To evaluate the influence of the combination of MOI and TOI on in vitro-replication of KHV-I, virus replication yield per cell was calculated based on the data obtained here The values ranged between ≤ 0.1 virus particles per cell for the highest tested MOI (0.012) and 1.82 virus particles per cell for MOI of 0.0007 and TOI of 30 000 cells/ cm2 (Table 2).
| TOI [cells/cm2] | MOI 0.0002 | MOI 0.0007 | MOI 0.003 | MOI 0.006 | MOI 0.012 |
|---|---|---|---|---|---|
| 15 000 | 0.48 | 0.59 | 0.45 | 0.40 | 0.02 |
| 30 000 | 0.59 | 1.82 | 0.43 | 0.59 | 0.02 |
| 60 000 | 0.22 | 0.74 | 0.18 | 0.74 | 0.26 |
| 75 000 | 0.19 | 0.49 | 0.15 | 0.80 | 0.14 |
Table 2: Maximum virus particles per cell obtained by inoculation of KHV-I at various MOI in CCB cells at various TOI. Calculated using Equation 2.
Discussion
As described above, the highest virus titers (≥ 105 TCID50/mL) were reached for the highest tested TOI (60 000 and 75 000 cells/cm2) and MOI between 0.0007 and 0.006, early after the inoculation (around 3 d p.i.). This was probably due to a high primary infection rate and therefore fast proliferation of secondary infection. Increasing MOI at a constant cell density at inoculation mostly resulted in an earlier peak of the virus titer (Figure 1). However, a higher amount of virus used for inoculation led to a faster infectivity loss during the infection course. Therefore, the TOH has to be appointed very carefully, since already a 2-days delay can result in harvesting at fivefold-lower virus titers. Furthermore, maximum virus titers obtained using the same cell density at inoculation but MOI of 0.0002 and 0.003 were lower in comparison to the MOI described above. Thus, leading to the conclusion that evaluation of the influence of MOI at constant TOI is quite difficult due to the high standard variation of the TCID50-assay.
When using low MOI (0.0002, 0.0007) in combination with all tested TOI, the maximum virus titers that were reached were often lower and generally observed later but also stayed stable for a longer time, resulting in a longer time frame for harvest. This might be due to the lower load of the viral particles at inoculation, leading to lower primary infection and postponed secondary infection. However, due to the lower virus load at inoculation and only slightly lower maximum virus titer, the maximum virus yield was higher (Table 1). Nevertheless, when using low cell densities, such as e.g. 15 000 cells/ cm2, for inoculation the maximum virus titer did not extend 104 TCID50/mL, regardless of applied MOI. Additionally, using high virus load for inoculation at low cell densities led to an early cell lysis preventing efficient replication of the virus.
The effects of variation of MOI, TOI and TOH were already analyzed in other systems such as: baculovirus (wild type and recombinant) in Sf9 insect cells, rabies virus and yellow fever virus in vero cells and KHV in various fish cell lines [25,31-34]. Comparable observations were made regarding the best time for harvest when inoculating with different virus load. Higher MOI often result in an early peak of the titer and a short time frame for harvest, whereas inoculation with low MOI often shows a shift of the titer peak to the time points later after inoculation [22,25,31,34,35]. Furthermore, it was reported for baculovirus that a high MOI leads to a synchronous, complete infection with a cessation of cell growth and can result in close to maximum productivity [32,36]. The results presented here suggest that this might also apply to KHV-I replication in CCB cells, however one has to be careful not to select too low TOI with too high MOI, since then cells, that are stressed not only due to the infection but also e.g. low concentrations of growth factors, might not be able to produce functional virus particles. Hence, resulting in very low or even depleted virus titers. In addition, when replicating the virus to obtain enough material for further investigation, higher effectivity (yield) can be reached by using lower MOI (Table 1). Therefore, both parameters: MOI and TOI must be carefully chosen so that too fast infection progress will not compromise reaching the highest virus titer.
Interestingly, our results showed that in general, the highest DNA load and the associated infectivity peak (TCID50/mL) occurred around the same time for KHV-I. This was already observed by Mletzko et al. [25] for KHV-T and indicates that qPCR results might give information about the earliest possible TOH. However, one has to be careful as while the viral DNA load often stays more or less stable afterwards, the corresponding infectivity can decrease even as soon as 1 day after reaching the peak. Obviously, in the later phase of replication the amount of non-infective particles or merely non-enveloped viral DNA rises considerably, for example due to the cell lysis. Furthermore, although isolated infection signs could be recorded via visual evaluation at the time point when the maximum titer occurred, the virus titer decreased again when severe infection was visible (Figure 3). Therefore, selection of the best time for harvest should not be based solely on the quantification of the viral DNA or visual investigation.
Mletzko et al. [25] already showed that infection of CCB with the KHV-T can result in maximum titers as high as 3.2 × 109 TCID50/ mL when using an intermediate TOI of 42 000 cells/cm2 and MOI of 0.003 Although various combinations of parameters were used for inoculation of KHV-I in the same cell line, comparably high titers were not reached within this work. This observation underlines the differences between the two isolates. In comparison to KHV-T, the replication of KHV-I was slower and not as efficient. Genome studies of three KHV isolates (-I, -USA, -Japan) concluded that these strains are highly conserved regarding nucleotide sequences, and show differences in only a small number of genes [30]. The majority of mutated genes encode for membrane glycoproteins, which might modulate the virus envelope and play a role in virus attachment and recognition by the cell. This could explain the differences in replication efficiency between KHV isolates. Though, investigation of three different KHV isolates from Europe and China (-FL, -GZ10, GZ-11) on protein level revealed that only few analyzed proteins were exclusively found in one of the isolates. Moreover, only one of these proteins was predicted to be a membrane protein and associated to the envelope of the virus. Regarding the other proteins, their localization could rarely be revealed [18,37].
Additionally, not reaching as high virus titers at lower MOI, as when using higher MOI with the same TOI, might be explained by the shift of the peak of the virus titer and thus nutrient limitation, increased concentration of metabolic products inhibiting virus proliferation, as well as by virus degrading processes resulting from enzymatic activity, due to cell lysis [22,25,34-36]. The latter might also explain the decrease of the virus titer after inoculation with TOI of 15 000 cells/cm2 and MOI of 0.012, since virus degradation counteracts virus replication. Furthermore, a high MOI is also often correlated with decreasing specific productivity and production of defective virus particles with negative influence regarding the virus yield [22]. Several studies recorded continuing cell growth post infection, especially when using low MOI, leading to a higher cell count and therefore, higher productivity [31,32,35,36]. Since KHV causes cell lysis, it is difficult to investigate cell growth after inoculation with the virus via actual cell counting. However, it is possible to assess the cell layer via microscopy in the early stages of viral infection, indicating that there is cell growth occurring post inoculation with KHV-I.
Experiments with baculovirus indicate that using high TOI (such as 4 × 106 cells/mL for baculovirus replication) leads in general to a decreased cell growth due to nutrient limitation, but that a supplementation with glutamine can result in higher virus titers [22]. Similar results regarding increase of the virus titer after glutamine supplementation were reported by Huang et al. [38] for influenza virus and MDCK cells. Nevertheless, whether this approach can also be applied to the KHV-I-CCB system needs clarification. This could be a valuable hint when using the cells for diagnostic purposes, where there is often a shortage of virus particles from environmental samples leading to slower replication progress. In such cases, supplementation of cells with nutrients might prevent limitation and lead to higher, observable virus titers even later after inoculation. However, for increasing the virus yield in general, it is more advantageous to infect with a low virus load, since the gain of the virus particles is higher, meaning one can reach similarly high titers by using less virus for inoculation.
Summary and Conclusion
In summary, our results show that the infection course of KHV-I in CCB cells is influenced by selected TOI and MOI. It is crucial to adjust both parameters to generate a maximum yield of virus particles and estimate the best time of harvest (TOH). Especially, the choice of TOI influences the optimum time point for harvest.
Although the work presented here showed that replication of KHV-I in CCB cells is not as efficient as in case of KHV-T, reproducible titers ≥ 5 × 104 TCID50/mL can also be reached for this isolate by careful adjustment of infection parameters. Therefore, in order to reach higher titers of KHV-I we recommend to use TOI of 60 000 to 75 000 cells/cm2 in combination with relatively low MOI values (~0.001).
Furthermore, based on the results presented here as well as on data obtained in our previous work and published by other research groups, it is clear that such studies should be performed for every KHV isolate, as they might behave very differently [25,27,29]. In addition, not only virus titers might not be high enough for further studies, but also the efficiency of cells for their virus replication could be falsely evaluated, due to disadvantageous choice of infection parameters. Finally, this work contributes to the better understanding of replication of various KHV isolates in CCB cells as well as provides new insights for general improvement of infection progresses for KHV in cell cultures. Nevertheless, further studies including further isolates of KHV are necessary.
Acknowledgements
This work was financially supported by the German Federal Ministry of Food and Agriculture (BMEL) through the Federal Office of Agriculture and Food (BLE), grant number 2815HS010.
References
- Waltzek TB, Kelley GO, Alfaro ME, Kurobe T, Davidson AJ, et al. (2009) Phylogenetic relationships in the family Alloherpesviridae. Dis Aquat Organ 84: 179-194.
- Hanson L, Dishon A, Kotler M (2011) Herpesviruses that infect fish. Viruses 3: 2160-2191.
- Haenen OLM, Way K, Bergmann SM, Ariel E (2004) The emergence of koi herpesvirus and its significance to European aquaculture. Bull Eur Assoc Fish Pathol 24: 293-307.
- Bergmann SM, Kempter J, Sadowski J, Fichtner D (2006) First detection, confirmation and isolation of koi herpesvirus (KHV) in cultured common carp (Cyprinus carpio L.) in Poland. Bull-Eur Assoc Fish Pathol 26: 97-104.
- Hedrick RP, Gilad O, Yun SC, McDowell TS, Waltzek TB, et al. (2005) Initial isolation and characterization of a herpes-like virus (KHV) from koi and common carp. Bull Fish Res Agency 2: V7.
- Dong C, Weng S, Li W, Li X, Yi Y, et al. (2011) Characterization of a new cell line from caudal fin of koi, Cyprinus carpio koi, and first isolation of cyprinid herpesvirus 3 in China. Virus Res 161: 140-149.
- Gray WL, Mullis L, LaPatra SE, Groff JM, Goodwin A (2002) Detection of koi herpesvirus DNA in tissues of infected fish. J Fish Dis 25: 171-178.
- Minamoto T, Honjo MN, Kawabata Z (2009) Seasonal distribution of Cyprinid Herpesvirus 3 in lake Biwa, Japan. Appl Environ Microbiol 75: 6900-6904.
- Eide KE, Miller-Morgan T, Heidel JR, Kent ML, Bildfell RJ, et al. (2011) Investigation of Koi Herpesvirus Latency in Koi. J Virol 85: 4954-4962.
- Xu J-R, Bently J, Beck L, Reed A, Miller-Morgan R, et al. (2013) Analysis of koi herpesvirus latency in wild common carp and ornamental koi in Oregon, USA. J Virol Methods 187: 372-379.
- Pikarsky E, Ronen A, Abramowitz J, Levavi-Sivan B, Hutoran M, et al. (2004) Pathogenesis of acute viral disease induced in fish by carp interstitial nephritis and gill necrosis Virus. J Virol 78: 9544-9551.
- Michel B, Fournier G, Lieffrig F, Costes B, Vanderplasschen A (2010a) Cyprinid herpesvirus 3. Emerg Infect Dis 16: 1835-1843.
- Costes B, Raj VS, Michel B, Fournier G, Thirion M, et al. (2009) The major portal of entry of Koi herpesvirus in Cyprinus carpio is the skin. J Virol 83: 2819-2830.
- Dishon A, Davidovich M, Ilouze M, Kotler M (2007) Persistence of cyprinid herpesvirus 3 in infected cultured carp cells. J Virol 81: 4828-4836.
- Gilad O, Yun S, Zagmutt-Vergara FJ, Leutenegger CM, Bercovier H, et al. (2004) Concentrations of a Koi herpesvirus (KHV) in tissues of experimentally-infected Cyprinus carpio koi as assessed by real-time TaqMan PCR. Dis Aquat Organ 60: 179-187.
- Brogden G, Adamek M, Proepsting MJ, Ulrich R, Naim HY, et al. (2015) Cholesterol-rich lipid rafts play an important role in the Cyprinid herpesvirus 3 replication cycle. Vet Microbiol 179: 204-212.
- Miwa S, Ito T, Sano M (2007) Morphogenesis of koi herpesvirus observed by electron microscopy. J Fish Dis 30: 715-722.
- Yi Y, Zhang H, Lee X, Weng S, He J, et al. (2014) Extracellular virion proteins of two Chinese CyHV-3/KHV isolates, and identification of two novel envelope proteins. Virus Res 191: 108-116.
- Mettenleiter TC (2004) Budding events in herpesvirus morphogenesis. Virus Res 106: 167-180.
- Bruss V, Ganem D (1991) The role of envelope proteins in hepatitis B virus assembly. Proc Natl Acad Sci U S A 88: 1059-1063.
- Cocquerel L, Wychowski C, Minner F, Penin F, Dubuisson J (2000) Charged residues in the transmembrane domains of hepatitis c virus glycoproteins play a major role in the processing, subcellular localization, and assembly of these envelope proteins. J Virol 74: 3623-3633.
- Maranga L, Brazão TF, Carrondo MJT (2003) Virus-like particle production at low multiplicities of infection with the baculovirus insect cell system: VLP Production at Low MOI. Biotechnol Bioeng 84: 245-253.
- Szilágyi, JF, Cunningham C (1991) Identification and characterization of a novel non-infectious herpes simplex virus-related particle. J Gen Virol 72: 661-668.
- Nielsen LK, Smyth GK, Greenfield PF (1992) Accuracy of the endpoint assay for virus titration. Cytotechnology 8: 231-236.
- Mletzko A, Amtmann A, Bergmann SM, Lee P, Christian J, et al. (2017) Inoculation of cyprinid herpesvirus 3 (CyHV-3) on common carp brain cells—influence of process parameters on virus yield. Vitro Cell Dev Biol - Anim 53: 579-585.
- Ronen A, Perelberg A, Abramowitz J, Hutoran M, Tinman S, et al. (2003) Efficient vaccine against the virus causing a lethal disease in cultured Cyprinus carpio. Vaccine 21: 4677-4684.
- Davidovich M, Dishon A, Ilouze M, Kotler M (2007) Susceptibility of cyprinid cultured cells to cyprinid herpesvirus 3. Arch Virol 152: 1541-1546.
- Hutoran M, Ronen A, Perelberg A, Ilouze M, Dishon A, et al. (2005) Description of an as yet unclassified DNA virus from diseased Cyprinus carpio species. J Virol 79: 1983-1991.
- Wang Y, Zeng W, Li Y, Liang H, Liu C, et al. (2015) Development and characterization of a cell line from the snout of koi (Cyprinus carpio L.) for detection of koi herpesvirus. Aquaculture 435: 310-317.
- Aoki T, Hirono I, Kurokawa K, Fukuda H, Nahary R, et al. (2007) Genome sequences of three Koi herpesvirus isolates representing the expanding distribution of an emerging disease threatening Koi and common carp worldwide. J Virol 81: 5058-5065.
- Radford KM, Cavegn C, Bertrand M, Bernard AR, Reid S, et al. (1997) The indirect effects of multiplicity of infection on baculovirus expressed proteins in insect cells: secreted and non-secreted products. Cytotechnology 24: 73-81.
- Freedman RB, Greenall C, Jenkins N, Tuite MF (1995) Protein folding in the secretory pathway of animal cells. In: Beuvery EC, Griffiths JB, Zeijlemaker WP (eds) Animal Cell Technology: Developments Towards the 21st Century. Springer, Dordrecht, The Netherlands, 371-376.
- Rourou S, Van der Ark A, Van der Velden T, Kallel H (2007) A microcarrier cell culture process for propagating rabies virus in Vero cells grown in a stirred bioreactor under fully animal component free conditions. Vaccine 25: 3879-3889.
- Souza MCO, Freire MS, Schulze EA, Gaspar LP, Castilho LR (2009) Production of yellow fever virus in microcarrier-based Vero cell cultures. Vaccine 27: 6420-6423.
- Baker K, Zheng YZ, Ried S, Greenfield PF (1993) Production of multisubunit particles for use as vaccines using the baculovirus expression vector system (BEVS). In: Animal Cell Technology: Basic & Applied Aspects. Springer, Dordrecht, The Netherlands, 521-534.
- Masoomi Dezfooli S, Tan WS, Tey BT, Ooi CW, Hussain SA (2016) Expression and purification of the matrix protein of Nipah virus in baculovirus insect cell system. Biotechnol Prog 32: 171-177.
- Michel B, Leroy B, Raj SV, Lieffrig F, Mast J, et al. (2010b) The genome of cyprinid herpesvirus 3 encodes 40 proteins incorporated in mature virions. J Gen Virol 91: 452-462.
- Huang D, Xia-Hou K, Liu XP, Zhao L, Fan L, et al. (2014) Rational design of medium supplementation strategy for improved influenza viruses production based on analyzing nutritional requirements of MDCK Cells. Vaccine 32: 7091-7097.
 Spanish
Spanish  Chinese
Chinese  Russian
Russian  German
German  French
French  Japanese
Japanese  Portuguese
Portuguese  Hindi
Hindi