Research Article, J Clin Exp Oncol Vol: 6 Issue: 7
Promoter Methylation and Loss of Expression of PTEN Gene in Breast Cancer Patients from Saudi Population
Mohammad Shabbir Alam1*, Ahmed Bin Ali Jerah1, Jalaluddin Mohammad Ashraf1, Kathamuthu Kumaresan1, Zaki Monawar Eisa2 and Nabil Tadros Mikhail2
1Department of Medical Laboratory Technology, College of Applied Medical Sciences, Jazan University, Saudi Arabia
2Department of Pathology, King Fahad Central Hospital, Abu Ariesh, Saudi Arabia
*Corresponding Author : Mohammad Shabbir Alam
Department of Medical Laboratory Technology, College of Applied Medical Sciences, Jazan University, Saudi Arabia
Tel: +966-535377920
E-mail: toshabbir@gmail.com
Received: September 25, 2017 Accepted: October 11, 2017 Published: October 18, 2017
Citation: Alam MS, Jerah ABA, Ashraf AM, Kumaresan K, Eisa ZM, et al. (2017) Promoter Methylation and Loss of Expression of PTEN Gene in Breast Cancer Patients from Saudi Population. J Clin Exp Oncol 6:6. doi: 10.4172/2324-9110.1000206
Abstract
Objective: PTEN, a tumor-suppressor gene, located on chromosome 10q23.3, is implicated in various types of cancer including breast cancer. The aim of this study is to investigate the promoter methylation, loss of expression and significance of PTEN gene in breast cancer and to determine the correlation between promoter methylation and gene expression.
Methods: Promoter methylation and loss of expression of PTEN gene were analyzed using methylation-specific PCR and immunohistochemical methods respectively. The chi square test is used to correlate the promoter methylation and gene expression with their clincopathologic parameters.
Results: We examined 53 breast cancer specimens and 10 normal tissues adjacent to tumor. The results showed a 58.5% promoter methylation in PTEN gene and none in normal tissue. PTEN methylation was observed in advanced stages III-IV (81.8%, 18 of 22, P=0.015) and higher grades G2-G3 (71.4%, 20 of 28, P=0.043) of disease. The correlation of PTEN methylation with clinical stage and tumor grade was found to be statistically significant. Nuclear PTEN expression was detected in 73.6% (39 of 53) cases of breast cancer and in the remaining 26.4% (14 of 53) cases expressional loss was observed. The loss of PTEN expression was observed in all normal tissues (10 of 10). The loss of PTEN expression was significantly correlated with patient’s age (P=0.028) and clinical stage (P = 0.029). The expressional loss was observed in 12 (38.7%) cases among 31 methylation positive cases, whereas among 22 methylation- negative cases, only 2 (9.1%) cases were seen as immunostaining negative with the statistically significant value (P=0.016).
Conclusion: Promoter methylation and loss of expression of PTEN gene occur frequently in breast cancer. Our results suggest that PTEN plays an important role in breast carcinogenesis.
Keywords: PTEN; Breast cancer; Methylation; Immunohistochemistry; Methylation-specific PCR, Estrogen receptor; Progesterone receptor
Abbreviations
ER: Estrogen Receptor; PR: Progesterone Receptor; WHO: World Health Organization; MSP: Methylation-specific PCR
Introduction
Breast cancer is the most common cancers among women worldwide [1]. It is also very common among Saudi females. Reports from different regions of Saudi Arabia demonstrate that majority of breast cancer patients are in the age group from 30 to 44 years [2]. However, this cancer is more common in older women above the age of 50 years worldwide but it is frequently diagnosed in younger women in Saudi Arabia.
PTEN is a tumor suppressor gene located at chromosome 10q23 [3,4]. It is a negative regulator of cell-survival signaling and has both lipid and protein phosphatase activity. Its major substrate is phosphatidylinositol-3,4,5-triphosphate (PIP-3), a direct product of phosphoinositol-3-kinase PI3K activity [5]. This substrate mediates growth factor-induced activation of intracellular signaling through serine-threonine kinase Akt, which promotes cell survival and proliferation. The Akt complex is activated through phosphorylation in actively proliferating cells. PTEN down-regulates the activity of Akt through dephosphorylation, thereby regulating phosphoinositol- 3kinase-triggered signaling negatively. Another study in breast cancer demonstrated that induction of apoptosis by low levels of PIP-3 and phosphorylated Akt is associated with high levels of PTEN [6]. PTEN gene is found to be mutated in a number of human malignancies such as melanoma [7], glioma [8], endometrial cancer [9], lung carcinoma [10], neuroblastic tumors [11], prostate cancer [12], cervical cancer [13] and breast cancer [14]. However, some investigators observed very low rate of somatic intragenic PTEN mutations and suggested that some other factors might be responsible for PTEN inactivation. Recently, the loss of PTEN expression has been discussed in many studies. In an immunohistochemical study, frequent loss or reduction of PTEN expression was observed in primary breast cancers [15]. Furthermore, the loss of PTEN expression due to promoter methylation of this gene is reported in cervical carcinoma, gastric cancer and non-small-cell lung cancer [13,16,17]. However, a little is known about the promoter methylation and its relation to loss of PTEN expression in breast cancer patients. Therefore, an effort was made in this study to investigate the PTEN promoter methylation and its correlation with the loss of PTEN expression in breast cancer.
Materials and Methods
Patients and tissue samples
Fifty three paraffin embedded blocks of tumor samples from breast cancer patients and 10 normal tissues adjacent to the tumors were collected between 2015 and 2017 from the archives of Department of Pathology, King Fahad Central Hospital, Abu Ariesh, Saudi Arabia. Clinical and pathological information, including age, tumor grade, clinical stage, lymph nodes metastasis and biomarkers, was also obtained. This study was approved by the ethics committee of King Fahad Central Hospital.
Sample preparation
Tissue sections (thickness, 5 μm) were cut from paraffin blocks. The sections were used to detect the PTEN expression in the breast carcinoma and normal breast tissues. Genomic DNA was also isolated from these sections using a ZR DNA FFPE Tissue Kit (Zymo- Research, USA) according to the manufacturer’s instructions. The purity and concentration of extracted DNA were analyzed by gel electrophoresis and ultraviolet spectrophotometry. DNA samples were frozen at ‑70ºC until use.
Analysis of promoter methylation of PTEN gene
DNA from tumor specimens as well as normal tissue was subjected to bisulfite modifications using an EZ DNA Methylation-Gold kit (Zymo Research, USA) as per procedure given in manufacturer’s protocol. The modified DNA was used as a template for methylation specific-PCR (MSP) as described previously [18]. Briefly, all samples were first amplified with primers specific for unmethylated DNA at the P15 locus to confirm the successful modification. All DNA samples were successfully amplified with the pair of primer, whereas unmodified DNA sample did not show amplification. The PCR was carried out in 25 μl volume containing 10 mM Tris-cl (pH 8.3), 50 mM KCl, 1.0 mM MgCl2, 100 μM dNTP, 0.5 μM of each primer, 1 unit of hot start Taq DNA polymerase (Thermo Fisher Scientific Inc.), and 25 ng bisulfite treated DNA. Primers for unmethylated p15 were 5’-GTGATGTGTTTGTATTTTGTGGTT-3’ and 5’-CCATACAATAACCAAACAACCAA- 3’, yielding a 154-bp product. Primers for methylated PTEN were 5’-TTCGTTCGTCGTCGTCGTATTT-3’ and 5’- GCCGCTTAACTCTAAACCGCAACCG- 3’, yielding a 206-bp product [19]. The PCR amplification conditions were followed as initial denaturation at 94°C for 5 min, followed by 32 cycles of 94ºC for 30s, 60°C (for unmethylated p15 gene) or 62°C (for methylated PTEN gene) for 30s and 72°C for 60s, followed by final extension at 72°C for 10 min. The PCR product (5 μl) was loaded on agarose gel (2%), staining was performed with ethidium bromide and observed under UV light. For positive control pUC19 bacterial strain DNA (methylation-negative) treated with SssI methyltransferase (Zymo Research, USA) followed by bisulfite treatment was used for methylated allele while the same DNA was used as a negative control for the unmethylated allele. A pUC19 DNA without pretreatment of SssI methyltransferase prior modification and a pUC19 DNA without any treatment were used as negative controls for the methylated allele.
Immunohistochemical (IHC) analysis
PTEN immunohistochemical staining was performed with an indirect biotin avidin system on the sections that were made from a representative block for each case. The slides were deparaffinized using xylene and graded ethyl alcohol, followed by rinsing with distilled water. Antigen retrieval was performed by boiling the slides in sodium citrate buffer, pH 6.0 in a microwave oven at 90ºC for 20 min. The sections were incubated in 0.3% hydrogen peroxide for 30 min to block the endogenous peroxidase activity. The slides were incubated with anti-PTEN antibody (Biogenic, San Ramon, CA, USA) overnight in refrigeration at 4ºC, subsequently incubated with a biotinylated universal secondary antibody and with an avidin horseradish peroxidase. After colour development with diaminobenzidine (DAB) nuclei were lightly counter stained with hematoxylin. For positive control stained slides (PTEN-positive) from patients with normal tissue were used in each set of experiment. For negative controls, the same procedure was followed with the exception of primary antibody. The staining was considered as either negative (-) or positive (+). The loss of PTEN expression was considered when the groups showing less than 5% tumor cell staining. It was assessed as negative. Anti-PTEN antibody (PTEN, [28H6]) is an IgG1 mouse monoclonal raised against human PTEN.
Statistical analysis
The statistical analysis was carried out with SPSS 18.0 software (Chicago, Illinois, USA) The Mantel-Haenszel χ2 test was used to determine the correlation of promoter methylation and PTEN gene expression with clincopathologic parameters. The relationship between promoter methylation with PTEN gene expression was also analyzed by χ2 test. Statistical differences were considered significant at P<0.05.
Results
Promoter methylation of PTEN gene
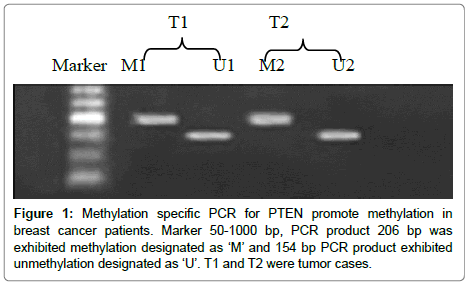
Promoter methylation of PTEN gene was analyzed in tumor as well as normal DNA from 53 breast cancer patients. Promoter methylation in PTEN gene was observed in 58.4% (31of 53) cases in the tumor DNA and none in the normal DNA (Figure 1). Table 1 shows the correlation between methylation and clincopathologic parameters.
| Characteristics | No. of cases (n=53) | Methylation status of PTEN | χ2 | P value | |
|---|---|---|---|---|---|
| Positive (%) | Negative (%) | ||||
| Age (years) ≤50 ˃50 |
32 21 |
17 (53.1) 14 (66.7) |
15 (46.9) 07 (33.3) |
0.957 | 0.328 |
| Tumor grade G1 G2-G3 |
25 28 |
11 (44.0) 20 (71.4) |
14 (56.0) 08 (28.6) |
4.092 | 0.043 |
| Lymph node metastasis No Yes |
28 25 |
19 (67.9) 12 (48.0) |
09 (32.1) 13 (52.0) |
2.145 | 0.143 |
| Stage I II III-IV |
12 19 22 |
05 (41.7) 08 (42.1) 18 (81.8) |
07 (58.3) 11 (57.9) 04 (18.2) |
8.431 | 0.015 |
| ER Positive Negative |
22 31 |
16 (72.7) 15 (48.4) |
06 (27.3) 16 (51.6) |
3.140 | 0.076 |
| PR Positive Negative |
25 28 |
12 (48.0) 19 (67.9) |
13 (52.0) 09 (32.1) |
2.145 | 0.143 |
| HER2 Positive Negative |
20 33 |
12 (60.0) 19 (57.6) |
08 (40.0) 14 (42.4) |
0.030 | 0.862 |
Table 1: Correlation of promoter methylation with clinicopathologic characteristics in breast cancer patients.
Among tumor specimen examined for promoter methylation according tumor grade, 44.0% (11 out of 25) cases were methylated in tumor grade G1 and 71.4% (20 out of 28) in grades GII-GIII. The relation between promoter methylation and tumor grade was found to be statistically significant (P=0.043). A statistically significant association between clinical stage and methylation was also observed (P=0.015) as 41.7% (5 of 12) cases were found to be methylated in clinical stage I, 42.1% (8 of 19) cases in stage II, and 81.8% (18 of 22) in stages III-IV. However, no such correlations were observed with patient’s age, tumor grade, and lymph node metastasis, ER, PR and HER2.
Analysis of PTEN gene expression
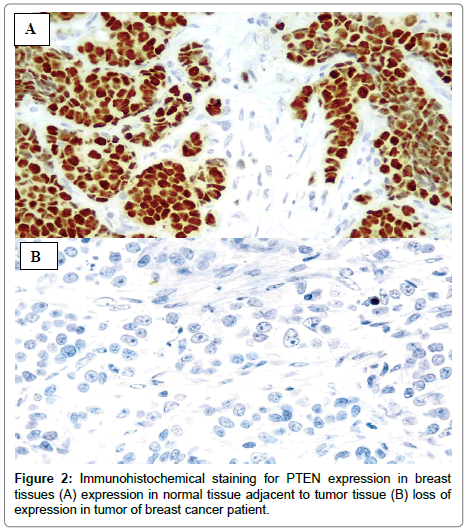
Immunohistochemical analysis of PTEN gene was performed in 53 cases of breast cancer and 10 normal tissues (adjacent to the tumors tissue) as a control. PTEN expression was detected in all 10 normal tissues and 39 of 53 (73.6%) cases in breast tumor whereas remaining 14 of 53 (26.4%) cases were observed as negative (Figure 2). The clinical and pathological characteristics were also correlated with PTEN expression (Table 2). The loss of expression was detected in 5 of 32 (15.6%) patients below the age of 50 years whereas 9 of 21 (42.9%) patients showed loss of PTEN expression above the age of 50 years. The loss of expression in PTEN gene was significantly correlated with the age (P=0.028). A significant relation was also observed between loss of PTEN expression and clinical stage (P=0.029) as loss of expression was detected in 1 of 12 (8.3%) cases in stage I, 9 of 19 (47.4%) in stage II and 4 0f 22 (18.2%) cases in stages III-IV. There were no such relations was noticed between tumor grade, lymph node metastasis, ER, PR and HER2.
| Characteristics | No. of cases n=53 |
PTEN gene expression | χ2 | P value | |
|---|---|---|---|---|---|
| Positive (%) | Negative (%) | ||||
| Age (years) ≤50 ˃50 |
32 21 |
27 (84.4) 12 (57.1) |
05 (15.6) 09 (42.9) |
4.837 | 0.028 |
| Tumor grade T1 T2-T3 |
25 28 |
19 (76.0) 20 (71.4) |
06 (24.0) 08 (28.6) |
0.142 | 0.706 |
| Lymph node metastasis No Yes |
28 25 |
21 (75.0) 18 (72.0) |
07 (25.0) 07 (28.0) |
0.061 | 0.804 |
| Stage I II III-IV |
12 19 22 |
11 (91.6) 10 (52.6) 18 (81.8) |
01 (08.3) 09 (47.4) 04 (18.2) |
7.077 | 0.029 |
| ER Positive Negative |
22 31 |
17 (77.3) 22 (70.9) |
05 (22.7) 09 (29.1) |
0.263 | 0.607 |
| PR Positive Negative |
25 28 |
20 (80.0) 19 (67.9) |
05 (20.0) 09 (32.1) |
1.001 | 0.317 |
| HER2 Positive Negative |
20 33 |
16 (80.0) 23 (69.7) |
04 (20.0) 10 (30.3) |
0.680 | 0.409 |
Table 2: Correlation of PTEN gene expressions with clinicopathologic characteristics of breast cancer patients.
The relation between promoter methylation and expression of PTEN gene
Correlation of promoter methylation with PTEN gene expression was determined in 53 cases of breast cancer. Thirty one cases were methylated and 22 cases were unmethylated at promoter region. Among 31 methylation-positive cases, loss of expression in PTEN gene was observed in 12 (38.7%) cases whereas among 22 methylation-negative cases, only 2 (9.1%) cases were seen as immunoreactions negative. The relation was found to be significant with their P value 0.016 (Table 3).
| PTEN Expression | Promoter methylation | χ2 | P value | |
|---|---|---|---|---|
| Methylated (%) | Unmethylated (%) | |||
| Positive Negative | 19 (61.3) 12 (38.7) | 20 (90.9) 2 (09.1) | 5.807 | 0.016 |
Table 3: Correlation of promoter methylation and PTEN gene expression in breast cancer patients.
Discussion and Conclusion
The role of PTEN identified as a novel tumor suppressor gene has been studied in a variety of cancers. Loss of heterozygosity and mutation of PTEN have been reported in melanoma, glioblastoma, kidney cancer, lung cancer, and breast cancer [20-22]. In particular, the mutation of PTEN appears with a frequency of up to 50% in endometrial cancer [23] compared with other cancers, including glioblastoma (28%), prostate cancer (30%), and melanoma (15%) [23]. In breast cancer, a 10-40% loss of heterozygosity (LOH) has been identified at the chromosome 10q23 region that contains the PTEN gene [25-27]. LOH at the 10q23 region is functionally associated with the occurrence of breast cancer. Although LOH at 10q23 is frequent, the somatic PTEN mutation rate is <5% and the majority of PTEN gene mutations occur in advanced and metastatic breast cancer [6].
In addition to gene mutation, epigenetic regulation, including promoter hyper methylation, has been demonstrated to alter tumor suppressor gene expression and contribute to tumorigenesis. As one of the most recurrent gene alterations, DNA methylation significantly affects chromosomal formation, gene expression and DNA replication [28,29]. CpG islands in the promoter or nearby regions are frequently methylated, leading to the silencing of gene transcription. Recent studies have also shown that CpG islands of the PTEN promoter are methylated in various human malignancies such as endometrial carcinoma [30], gastric [16], non-small-cell lung carcinoma [17], cervical cancer [13,31], including breast cancer [14,32]. The methylation status of the PTEN promoter CpG Island was analyzed in the present study. Promoter methylation was seen in 58.5% (31 of 53) cases of breast carcinoma. Clinicopathologic parameters were correlated with the PTEN promoter methylation. A significant correlation was observed between PTEN methylation and, tumor grade and tumor stage. A high methylation rate (71.4%) was occurred in G2-G3 grades as compared to lower grade G1 (44.2%). The advanced stages III-IV of tumors was also exhibited the high frequency of promoter methylation when compared to stages I and II (Table 3). These findings suggest that promoter methylation of PTEN gene is associated with advanced stage and higher grade of the disease.
The loss of PTEN expression is thought to play an important role in tumor cell proliferation and metastasis; due to lack of control of the signaling pathways that mediate cellular processes such as apoptosis and migration [33,34]. Therefore, we performed immunohistochemistry to evaluate PTEN expression in breast cancer. The loss of PTEN expression was found in 26.4% (14 of 53) cases of breast carcinoma. These findings suggest that loss of PTEN expression play a role in carcinogenesis of breast. A significant correlation was observed between clinical stage and expressional loss of PTEN (Table 2). Statistically significant association with respect to loss of PTEN expression with age could only be noticed among the patients of more than 50 years of age. The expressional loss of PTEN was also observed in advanced stages III-IV of tumor as compared to stage I and Stage II (Table 2). Thus, the present study shows that clinical stage and patient’s age are important factors in determining the level of PTEN expression and eventually influencing the occurrence of breast cancer.
Some previous studies have also shown on association of PTEN promoter methylation with its reduced expression in breast cancer [14,35,36]. A correlation between the occurrences of PTEN promoter methylation with its reduced expression was also made. Twelve of 31 (38.7%) cases of breast cancer showing promoter methylation also revealed the loss of PTEN expression, whereas 9.1% (2 of 22) breast carcinoma showing no methylation exhibited the expressional loss of PTEN. Promoter methylation of PTEN was significantly correlated with the loss of PTEN expression and showed a significant relationship (P=0.016). These results indicate that reduced PTEN expression may be due to PTEN promoter methylation. Thus, the present study is the first report of its kind from Saudi population to show the association between promoter methylation and PTEN gene silencing in breast cancer.
In conclusion, our findings demonstrate that promoter methylation and loss of PTEN expression occur frequently in breast cancer patients. Promoter methylation of PTEN presents a major alternative mechanism of gene silencing and appears to play an important role in breast carcinogenesis.
Acknowledgment
The research was supported by Grants (No. 36/6/5021) from the Deanship of Scientific Research, Jazan University, Jazan, Kingdom of Saudi Arabia. The authors acknowledge Dr. Ahmed Abdul Haque, Dean, Deanship of Scientific Research, Dr. Yahya Hasan Hobani, Dean, College of Applied Medical Sciences and Prof. Anil Kumar Bidwai for their help and support.
Conflict of Interest
The authors report no conflict or interest in this work.
References
- WHO (2007) Fight against Cancer: Strategies that prevent, cure and care, Geneva, Switzerland.
- Saudi Cancer Registry (2007) Saudi Arabia Cancer Incidence and Survival Report 2007, Riyadh, Saudi Arabia.
- Li J, Yen C, Liaw D, Podsypanina K, Bose S, et al. (1997) PTEN, a putative protein tyrosine phosphatase gene mutated in human brain, breast, and prostate cancer. Science 275: 1943-1947.
- Steck PA, Pershouse MA, Jasser SA, Yung WK, Lin H, et al. ( 1997) Identification of a candidate tumour suppressor gene, MMAC1, at chromosome 10q23.3 that is mutated in multiple advanced cancers. Nat Genet 15: 356-362.
- Ali IU, Schriml LM, Dean M (1999) Mutational spectra of PTEN/MMAC1: a tumor suppressor with lipid phosphatase activity. J Natl Cancer Inst 91: 1922-1932.
- Lu Y, Lin YZ, LaPushin R, Cuevas B, Fang X, et al. (1999) The PTEN/MMAC1/TEP tumor suppressor gene decreases cell growth and induces apoptosis and anoikis in breast cancer cells. Oncogene 18: 7034‑7045.
- Guldberg P, Straten P, Birck A, Ahrenkiel V, Kirkin AF, et al. (1997) Disruption of the MMAC1/PTEN gene by deletion or mutation is a frequent event in malignant melanoma. Cancer Res 57: 3660-3663.
- Rasheed BK, Stenzel TT, McLendon RE, Parsons R, Friedman AH, e al. (1997) PTEN gene mutations are seen in high-grade but not in low-grade gliomas. Cancer Res 57: 4187-4190.
- Aneta JJ, Bo-zena K, Cyprian G, Urszula N (2010) TP53 Mutations in Endometrial Cancers: Relation to PTEN Gene Defects. Int J Gynecol Cancer 20: 196-202.
- Zhang L, Liu T, Liu H, Gao J, Gu C (2000) Loss and inactivation of PTEN/MMAC1/TEP1 gene in lung cancer. Zhonghua Bing Li Xue Za Zhi 29: 85-88.
- Izycka-Swieszewska E, Brzeskwiniewicz M, Wozniak A, Drozynska E, Grajkowska W, et al. (2010) EGFR, PIK3CA and PTEN gene status and their protein product expression in neuroblastic tumours. Folia Neuropathol 48: 238-245.
- Reid AHM, Attard G, Ambroisine L, Fisher G, Kovacs G, et al (2010) Molecular characterisation of ERG, ETV1 and PTEN gene loci identifies patients at lowand high risk of death from prostate cancer. Br J Cancer 102: 678-684.
- Rizvi MMA, Alam MS, Ali A, Mehdi SJ, Batra S, et al. (2011) Aberrant promoter methylation and inactivation of PTEN gene in cervical carcinoma from Indian population. J Cancer Res Clin Oncol 137: 1255-1262.
- Zhang HY, Liang F, Jia ZL, Song ST, Jiang ZF (2013) PTENmutation, methylation and expression in breast cancer patients. Oncol Lett 6: 161-168.
- Perren A, Weng LP, Boag AH, Ziebold U, Thakore K, et al. (1999) Immunohistochemical evidence of loss of PTEN expression in primary ductal adenocarcinomas of the breast. Am J Pathol 155: 1253-1260.
- Kang YH, Lee HS, Kim WH (2002) Promoter methylation and silencing of PTEN in gastric carcinoma. Lab Invest 82: 285-291.
- Soria JC, Lee HY, Lee JI, Wang L, Issa JP, et al. (2002) Lack of PTEN expression in non-small cell lung cancer could be related to promoter methylation. Clin Cancer Res 8: 1178-1184.
- Cheung TH, Lo WK, Yim SF, Chan LK, Jeung MS, et al. (2004) Epigenetic and genetic alteration of PTEN in cervical neoplasm. Gynecol Oncol 93: 621-627.
- Herman JG, Graff JR, Myohanen S, Nelkin BD, Baylin SB (1996) Methylation-specific PCR. A novel PCR assay for methylation status of CpG islands. Proc Natl Acad Sci 93: 9821-9826.
- Rhei E, Kang L, Bogomolniy F, Federici MG, Borgen PI, et al. (1997) Mutation analysis of the putative tumor suppressor gene PTEN/MMAC1 in primary breast carcinomas. Cancer Res 57: 3657-3659.
- Risinger JI, Hayes K, Maxwell GL, Carney ME, Dodge RK, et al. (1998) PTENmutation in endometrial cancers is associated with favorable clinical and pathologic characteristics. Clin Cancer Res 4: 3005-3010.
- Teng DH, Hu R, Lin H, Davis T, Iliev D, et al. (1997) MMAC1/PTENmutations in primary tumor specimens and tumor cell lines. Cancer Res 57: 5221-5225.
- Tashiro H, Blazes MS, Wu R, Cho KR, Bose S, et al. (1997) Mutations in PTENare frequent in endometrial carcinoma but rare in other common gynecological malignancies. Cancer Res 57: 3935-3940.
- Dahia PLM (2000) PTEN, a unique tumor suppressor gene. Endocr Relat Cancer 7: 115-129.
- Marsh DJ, Coulon V, Lunetta KL, Rocca‑Serra P, Dahia PL, et al. (1998) Mutation spectrum and genotype‑phenotype analyses in Cowden disease and Bannayan‑Zonana syndrome, two hamartoma syndromes with germline PTEN mutation. Hum Mol Genet 7: 507‑515.
- Freihoff D, Kempe A, Beste B, Wappenschmidt B, Kreyer E, et al. (1999) Exclusion of a major role for the PTEN tumour‑suppressor gene in breast carcinomas. Br J Cancer 79: 754‑758.
- Bose S, Wang SI, Terry MB, Hibshoosh H, Parsons R (1998) Allelic loss of chromosome 10q23 is associated with tumor progression in breast carcinomas. Oncogene 17: 123‑127.
- Keshet I, Lieman‑Hurwitz J, Cedar H (1986) DNA methylation affects the formation of active chromatin. Cell 44: 535‑543.
- Antequera F (2003) Structure, function and evolution of CpG island promoters. Cell Mol Life Sci 60: 1647‑1658.
- Salvesen HB, MacDonald N, Ryan A, Jacobs IJ, Lynch ED, et al. (2001) PTEN methylation is associated with advanced stage and microsatellite instability in endometrial carcinoma. Int J Cancer 91: 22-26.
- Yang HJ, Liu SW, Wang Y, Tsang CK, Ngan YS (2006) Differential DNA methylation profiles in gynecological cancers and correlation with clinic-pathological data. BMC Cancer 6: 212.
- Luo S, Chen J, Mo X (2016) The association of PTEN hypermethylation and breast cancer: a meta analysis. Onco Targets and Therapy 9: 5643-5650.
- Huang J, Kontos CD (2002) PTEN modulates vascular endothelial growth factor-mediated signaling and angiogenic effects. J Biol Chem 277: 10760-10766.
- Kandel ES, Skeen J, Majewski N, Di Cristofano A, Pandolfi PP, et al. (2002) Activation of Akt/protein kinase B overcomes a G (2)/m cell cycle checkpoint induced by DNA damage. Mol Cell Biol 22: 7831-7841.
- Siddiqui S, Akhtar N, Deo SVS, Shukla NK, Hussain SA (2016) A study on promoter methylation of PTEN in sporadic breast cancer patients from north India. Breast Cancer 23: 922-931.
- Lu YM, Cheng F, Teng LS (2016) The association between phosphatase and tensin homologhypermethylation and patients with breast cancer, a meta-analysis and literature review. Sci Rep 6: 32723.
 Spanish
Spanish  Chinese
Chinese  Russian
Russian  German
German  French
French  Japanese
Japanese  Portuguese
Portuguese  Hindi
Hindi