Review Article, J Mol Biol Methods Vol: 1 Issue: 1
Chloroplast DNA: A Promising Source of Information for Plant Phylogeny and Traceability
Freitas AS1, da Anunciação RR2, D’Oliveira-Matielo CB3 and Stefenon VM3*
1Centro Interdisciplinar de Pesquisas em Biotecnologia, Universidade Federal do Pampa, Campus São Gabriel, São Gabriel, Rio Grande do Sul, Brasil
2Núcleo de Estudos da Vegetação Antártica, Universidade Federal do Pampa, Campus São Gabriel, São Gabriel, Rio Grande do Sul, Brasil
3Núcleo de Ecologia Molecular e Micropropagação de Plantas, Universidade Federal do Pampa, Campus São Gabriel, São Gabriel, Rio Grande do Sul, Brasil
*Corresponding author : Valdir Marcos Stefenon
Núcleo de Ecologia Molecular e Micropropagação de Plantas, Universidade Federal do Pampa, Campus São Gabriel, São Gabriel, Rio Grande do Sul, Brasil
Tel: +55 55 3237 0858
E-mail: valdirstefenon@unipampa.edu.br
Financial support: CAPES Pro-Forense Consortium and CNPq
Received: December 23, 2017 Accepted: February 21, 2018 Published: March 01, 2018
Citation: Freitas AS, da Anunciação RR, D’Oliveira-Matielo CB, Stefenon VM (2018) Chloroplast DNA: A Promising Source of Information for Plant Phylogeny and Traceability. J Mol Biol Methods 1:1.
Abstract
Chloroplasts are organelles with a specific DNA (cpDNA), which are responsible by the energetic metabolism in eukaryotic cells from plants and algae. Their key role is to perform the photosynthesis, when the cell converts sunlight and carbon dioxide into glucoses (=energy) and oxygen. Studies aiming to understand how chloroplast works are important to highlight details about plant evolution. The new approaches in molecular biology have raised the knowledge about the chloroplast genome characteristics and opened a wide range of possible applications for the increased background. In this review, we discuss the use of chloroplast DNA for plant phylogeny and traceability. Considering the increasing need for food safety, identification of gene flow from genetically modified plants and for better tools to police investigation, many works have demonstrated the viability of using cpDNA for this purposes, either for phylogeny of land plants or traceability of food, genetically modified plants and illicit drugs.
Keywords: cpDNA; Food safety; Genetically modified plants; Traceability
Introduction
Chloroplasts are organelles responsible for the energetic metabolism in autotrophic eukaryotic cells that set up the gathering and conversion of light and carbon dioxide into glucoses and oxygen. They are found in plants and algae, corresponding to a center of energy production for the cells [1]. Actually, these organelles are distinguished from other types of plastids by the presence of chlorophyll, being most concentrated in the leaf mesophyll [2].
The key role of chloroplasts is to conduct and perform the photosynthesis, process in which the pigment chlorophyll captures the energy from sunlight, and converts and stores it in ATP and NADPH molecules while freeing oxygen from water. Subsequently, ATP and NADPH are used to synthetize organic molecules from carbon dioxide [3]. Leaf guard cells of most plant species contain chloroplasts that possess an active electron transport chain, being capable of fixing carbon dioxide and play a fundamental role in stomatal opening and closing [2].
The origin of the chloroplasts is correlated to an endosymbiosis event in which a eukaryotic cell encompassed a photosynthetic bacterium and both eukaryotic cell and bacteria adapted themselves to live in symbiosis [1,4]. This theory supports and explains why chloroplasts have their own genome and transcription machinery, as well its specific genetic code, not linked to the nuclear genome.
On this context, studies that goal to elucidate mechanisms of regulation of chloroplast metabolism are very important to understand how this organelle performs its biological activities and guarantee the adaptive success of plants. In this direction, new approaches in molecular biology – such as the Next Generation Sequencing (NGS) – have increased quickly the knowledge about genome sequence and functions in a wide range of organisms from bacteria to humans and complex plants [5].
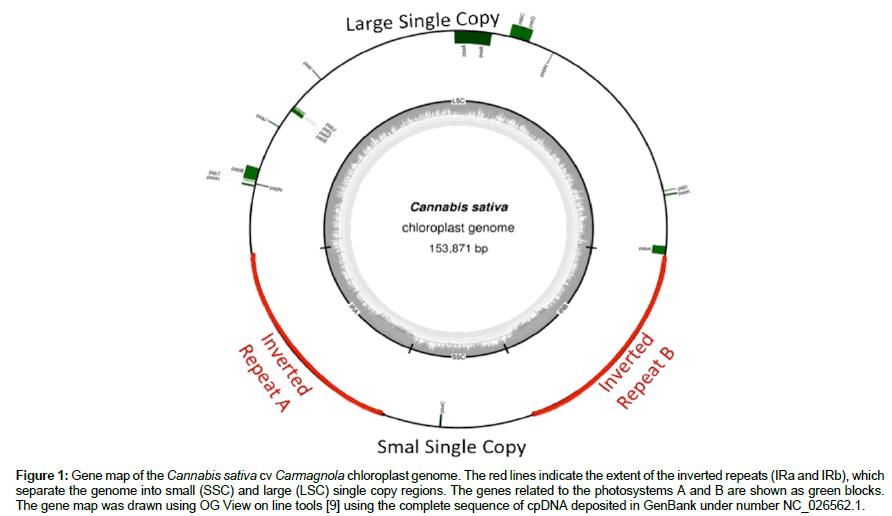
The first chloroplasts genome study was officially reported in 1986 with the publishing of the complete tobacco (Nicotiana tabacum) chloroplast genome [6]. The chloroplast DNA, often abbreviated as cpDNA, is composed by a single large circular DNA molecule; typically 120,000-170,000 base pairs long (Figure 1). They can have a contour length of around 30-60 micrometers, and have a mass of about 80-130 million daltons [7]. Chloroplasts sequences studies – as well as mitochondrial sequences – have taken prominence in works of plant evolution, phylogeny and traceability due to its small length, simple structure, maternal inheritance characters and conserved sequences [8].
Figure 1: Gene map of the Cannabis sativa cv Carmagnola chloroplast genome. The red lines indicate the extent of the inverted repeats (IRa and IRb), which separate the genome into small (SSC) and large (LSC) single copy regions. The genes related to the photosystems A and B are shown as green blocks. The gene map was drawn using OG View on line tools [9] using the complete sequence of cpDNA deposited in GenBank under number NC_026562.1.
The efficacy of DNA in relation to protein analyzes are already recognized as being applied to a variety of matrices and easily extracted with low biomass [9,10]. Currently, a number of existing biomarkers models (RAPDs, AFLPs, SSRs, SNPs and SCARs) are sources of potential DNA fingerprinting generators, creating a specific identification for each organism [11], and can identify the differences between organisms at different taxonomic levels, like genera or, more specifically, differentiating distinct varieties within the same species.
In this review, we present and discuss the main biotechnological resources for phylogeny and traceability of plants based on chloroplast DNA sequences analyses.
Phylogeny of plants
Noncoding sequences from the chloroplast genome are a primary source of data for molecular systematic, phylogeographic, and population genetic studies of plants. Even though still relatively little is known about levels of variation among different regions of the chloroplast genome, the facility of working with and the low cost for sequencing have increased the interest by this approach [7].
Due to previous related characteristics, genes presents in chloroplast genomes have shown a great potential for phylogenetic markers, such as performed by many works [7,12,13]. These efforts have contributed for a new comprehension of genetic plant relationship. Clearly, a phylogenomic approach based on complete chloroplast genomes has the potential to resolve some important taxonomic inconsistencies still discussed by taxonomists. As an example, we built a phylogenomic tree based on the complete sequences of chloroplast genomes from 19 species and clearly resolved the major taxonomic groups (Figure 2). Even the polytomy observed for the two species of Bryophyta may be an insight to review the origin of this group.
Traceability of food
Concern about food safety as well as its authenticity, quality and legitimacy has increased due to bioterrorism assessment and the emerging implications of outbreaks of foodborne diseases [13,14]. Techniques based on genetic markers and PCR assay promote the growing required safety in products that possesses in its composition any vegetal base [11,15,16].
Genetic traceability arises as a way to include information about the origin, materials or ingredients present in plant products, as well as to determinate the genetic makeup of the products [13]. These approaches allow authenticating and investigating the veracity of the kind of plant used in product design even as in various processing phases [17].
Currently, studies aiming this concern have been carried out [11,18]. Some researchers have already sought to fill gaps in the traceability of industrialized products and products that are not directly based on the use of plants. Making traceability a reality in the trophic chain, as well as highly processed products [15,19].
Traceability of genetically modified plants (GMPs)
Although intense research has been performed about the safety for the human health and/or for the environment – and no significant hazards connected to its use has been detected –, genetically modified plants (also know as transgenic plants) are a still polemic point of discussion [20]. Authors as Nicolia and colleagues show the current concern of the academic society regarding transgenic plants and their relation to the environment [21]. The need for security and the fear of topics such as gene flow, low biodiversity and indicative of tumors incidence [22] are points that generate doubts about the real security of implementation of this plants, even though nothing was proved and methods used in some studies have been criticized [23,24].
An interesting point of view is the use of genetic markers in the genome of GMPs to evaluate the dissemination of modified DNA. As many transformations of plants are performed in the chloroplast genome, such as for production of interest proteins [25,26], cpDNA markers are a very viable alternative to trace the dissemination of genetic material along a population of plants and prevent possible undesirable events.
The use of traceability is already performed in the European Union [27] and in the USA. However, due to the problems already mentioned, there is still no homogeneity regarding the marking of these products or the methods of analysis [20], demonstrating that many efforts are still necessary to a viable and confident method for GMP’s traceability. Although no standard method exists, the scenario is very optimistic.
Traceability of illicit drugs
Illicit drugs are a problem of public health in the world, mainly in developing countries [28]. In a context of scarcity of information about this theme, to know the geographical origin from where the drugs consumed by people came from becomes very important information for police and health agents. Trace the routes of trafficking may help competent authorities to elucidate and combat crimes related to drugs [29]. On the other hand, traceability of plants such as Cannabis sativa and Erythroxylum coca may also provide genetic information that differentiates plants cultivated for medicinal use, from plants cultivated for illicit use.
The prospection and application of cpDNA markers from plants as marijuana (Cannabis sativa L.) are a possible alternative to better map the origin of seized plants and drugs. No definitive study about this theme has been published yet, but some groups have tried to determinate the relationships between different varieties of C. sativa. Gonçalves [30], for example, observed distinctions in genes from different Cannabis samples seized by the Brazilian Federal Police in this country, while a panel of nuclear SSR markers was tested for drugs seizures in Australia and USA [31-33].
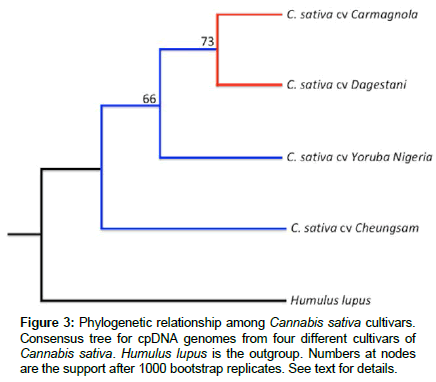
The usefulness of chloroplast genome sequences for forensic proposes was recently demonstrated by our group. We found significant differences among C. sativa cultivars based on plastidial DNA (Figure 3), showing that this approach based on cpDNA is a viable and reproducible tool for traceability of this species. Two cultivars designated for medicinal use due to low cannabidiol content (Carmagnola and Dagestani) grouped together, while cultivars Yoruba Nigeria and Cheungsam (which present high cannabidiol content and are cultivated and commercialized as drug) are in different clusters (Figure 3). This result suggests that even though the genes related to the cannabidiol are located in the nuclear genome, the selection of plants can also be recognized through the chloroplast genome.
Conclusions
Since the beginning of the novel sequencing techniques, much information about cpDNA, as well as its intraspecific relationships, offered a great contribution to the scientific world. In the last 30 years, such research allowed the development of technologies and tools that are rapidly advancing and improving global security and traceability in plant-based products as well as uncovering questions about the plant evolution and adaptation.
Based on these data, we show that using complete chloroplast genome arises as a promising tool for classification and differentiation of plants. More researches as well as the sequencing of more chloroplasts genomes are needed to a better understand about phylogenetic relationships among plants and for the establishment of standard protocols for DNA traceability. Nonetheless, existing studies already show an exciting scenario.
Acknowledgement
Authors thank to UNIPAMPA by general logistic, to CAPES (Pro-Forense Consortium) by a scholarship and financial support for CBD’O-M and VMS and to CNPq by a scholarship for ASF and RRA.
References
- Chan CX, Bhattacharya D (2010) The origin of plastids. Nat Educ 3: 84-91.
- Barton KA, Wozny MR, Mathur N, Jaipargas E-A, Mathur J (2017) Chloroplast behaviour and interactions with other organelles in Arabidopsis thaliana pavement cells. J Cell Sci 131: jcs202275.
- Alberts B, Johnson A, Lewis J, Morgan D, Raff M, et al. (2017) Biologia Molecular da Célula Artmed (6th edtn) Porto Alegre, Brazil.
- Sánchez-Baracaldo P, Raven JA, Pisani D, Knoll AH (2017) Early photosynthetic eukaryotes inhabited low-salinity habitats. Proc Natl Acad Sci U S A 114: E7737-E7745.
- Schuster SC (2007) Next-generation sequencing transforms today’s biology. Nat Methods 5: 16-18.
- Shinozaki K, Ohme M, Tanaka M, Wakasugi T, Hayashida N, et al. (1986) The complete nucleotide sequence of the tobacco chloroplast genome: its gene organization and expression. EMBO J 5: 2043-2049.
- Shaw J, Lickey EB, Schilling EE, Small RL (2007) Comparison of whole chloroplast genome sequences to choose noncoding regions for phylogenetic studies in angiosperms: the tortoise and the hare III. Am J Bot 94: 275-288.
- Xin T, Dezhu L (2002) Application of DNA sequences in plant phylogenetic study. Acta Bot Yunnanica 24: 170-184.
- Lohse M, Drechsel O, Kahlau S, Bock R (2013) OrganellarGenomeDRAW- a suite of tools for generating physical maps of plastid and mitochondrial genomes and visualizing expression data sets. Nucleic Acids Res 41: W575-W581.
- Galimberti A, De Mattia F, Losa A, Bruni I, Federici S, et al. (2013) DNA barcoding as a new tool for food traceability. Food Res Int 50: 55-63.
- Agrimonti C, Vietina M, Pafundo S, Marmiroli N (2011) The use of food genomics to ensure the traceability of olive oil. Trends Food Sci Technol 22: 237-244.
- Shaw J, Lickey EB, Beck JT, Farmer SB, Liu W, et al. (2005) The tortoise and the hare II: relative utility of 21 noncoding chloroplast DNA sequences for phylogenetic analysis. Am J Bot 92: 142-166.
- Opara LU (2003) Traceability in agriculture and food supply chain: a review of basic concepts, technological implications, and future prospects. J Food Agric Environ 1: 101-106.
- APG (2009) An update of the Angiosperm Phylogeny Group classification for the orders and families of flowering plants: APG III. Bot J Linn Soc 161: 105-121.
- Martellossi C, Taylor EJ, Lee D, Graziosi G, Donini P (2005) DNA extraction and analysis from processed coffee beans. J Agric Food Chem 53: 8432-8436.
- Wambulwa MC, Meegahakumbura MK, Kamunya S, Muchugi A, Möller M, et al. (2016) Insights into the genetic relationships and breeding patterns of the African tea germplasm based on nSSR Markers and cpDNA Sequences. Front Plant Sci 7: 1244.
- Doveri S, Lee D (2007) Development of sensitive crop-specific polymerase chain reaction assays using 5s dna: applications in food traceability. J Agric Food Chem 55: 4640-4644.
- García-González DL, Aparicio R (2010) Research in olive oil: challenges for the near future. J Agric Food Chem 58: 12569-12577.
- Ponzoni E, Mastromauro F, Gianì S, Breviario D (2009) Traceability of plant diet contents in raw cow milk samples. Nutrients 1: 251-262.
- Klümper W, Qaim M (2014) A meta-analysis of the impacts of genetically modified crops. PLoS One 9: e111629.
- Nicolia A, Manzo A, Veronesi F, Rosellini D (2014) An overview of the last 10 years of genetically engineered crop safety research. Crit Rev Biotechnol 34: 77-88.
- Seralini GE, Mesnage R, Clair E, Gress S, de Vendomois J, et al. (2011) Genetically modified crops safety assessments: present limits and possible improvements. Environ Sci Eur 23: 10.
- Tien DL, Huy HL (2013) Comments on "Long term toxicity of a roundup herbicide and a roundup-tolerant genetically modified maize". Food Chem Toxicol 53: 443-444.
- Tribe D (2013) Letter to the editor. Food Chem Toxicol 53: 467-472.
- Avila EM, Day A (2014) Stable Plastid Transformation of Petunia. In: Maliga P (eds) Chloroplast Biotechnology: Methods Mol Biol 1132: 277-293.
- Economou C, Wannathong T, Szaub J, Purton S (2014) A simple, low-cost method for chloroplast transformation of the green alga chlamydomonas reinhardtii. In: Maliga P (eds) Chloroplast Biotechnology: Methods Mol Biol 1132: 401-411.
- Naegeli H, Birch AN, Casacuberta J, De Schrijver A, Gralak MA, et al. (2017) Guidance for the risk assessment of the presence at low level of genetically modified plant material in imported food and feed under Regulation (EC) No 1829/2003. EFSA J 15: e05048.
- Johnston LD, O’Malley PM, Bachman JG, Schulenberg JE (2013) Demographic subgroup trends among adolescents for fifty-one classes of licit and illicit drugs, 1975-2012. University of Michigan, USA.
- Williams J, Banta-Green C, Burgard D (2017) The need for better marijuana sales data. Addiction 112: 2179-2180.
- Gonçalves FC (2015) Molecular identification and phylogenetic analysis of Cannabis samples seized by the Civil Police of the State of Espírito Santo. Federal University of Espirito Santo, Brazil.
- Gilmore S, Peakall R, Robertson J (2003) Short tandem repeat (STR) DNA markers are hypervariable and informative in Cannabis sativa: implications for forensic investigations. Forensic Sci Int 131: 65-74.
- Howard C, Gilmore S, Robertson J, Peakall R (2009) A Cannabis sativa STR genotype database for Australian seizures: forensic applications and limitations. J Forensic Sci 54: 556-563.
- Köhnemann S, Nedele J, Schwotzer D, Morzfeld J, Pfeiffer H (2012) The validation of a 15 STR multiplex PCR for Cannabis species. Int J Legal Med 126: 601-606.
 Spanish
Spanish  Chinese
Chinese  Russian
Russian  German
German  French
French  Japanese
Japanese  Portuguese
Portuguese  Hindi
Hindi