Research Article, J Appl Bioinforma Comput Biol Vol: 7 Issue: 2
Use of Chemo-Informatics to Identify Molecular Descriptors of Auxins, Cytokinins and Gibberellins
Ivan Andújar1, Daviel Gomez2, Lianny Perez3 and Jose Carlos Lorenzo2*
1Laboratory for Plant Cell and Tissue Culture, Bioplant Center, University of Ciego de Avila, Ciego de Avila, 69450, Cuba
2Laboratory for Plant Breeding & Conservation of Genetic Resources, Bioplant Center, University of Ciego de Avila, Ciego de Avila, 69450, Cuba
3 Laboratory for Metabolic Engineering, Bioplant Center, University of Ciego de Avila, Ciego de Avila, 69450, Cuba
*Corresponding Author : Jose Carlos Lorenzo
Laboratory for Plant Breeding & Conservation of Genetic Resources, Bioplant Center, University of Ciego de Avila, Ciego de Avila, 69450, Cuba
E-mail: jclorenzo@bioplantas.cu
Received: May 05, 2018 Accepted: May 25, 2018 Published: June 01, 2018
Citation: Andújar I, Gomez D, Perez L, Lorenzo JC (2018) Use of Chemo-Informatics to Identify Molecular Descriptors of Auxins, Cytokinins and Gibberellins. J Appl Bioinforma Comput Biol 7:2. doi: 10.4172/2329-9533.1000151
Abstract
We have identified those molecular descriptors differentiating four auxins, four cytokinins and four gibberellins. DRAGON software (version 5.5, 2007) and CambridgeSoft ChemOffice (version 12, 2010) including ChemDraw and Chem3D were used to calculate 212 molecular descriptors. Only 49 descriptors showed statistically significant differences among auxins, cytokinins and gibberellins. Of them, the most important differences can be described as follows. Gibberellins contain terminal tertiary C (sp3), terminal quaternary C (sp3), ring secondary C (sp3), ring tertiary C (sp3), and ring quaternary C (sp3) that are not present either in cytokinins or auxins. Gibberellins are also relatively rich in terminal secondary C (sp3) and 10-membered rings which are absent in cytokinins. Cytokinins have 10 times more nitrogen atoms than auxins but this atom is not present in gibberellins. Auxins have 10 times more substituted benzene C (sp2) and 5 times more benzene-like rings than cytokinins but these structures are not in gibberellins. Regarding the numbers of unsubstituted benzene C (sp2), auxins average 4.50, cytokinins 1.25 but they are absent in gibberellins. A dendogram was generated using data of those molecular descriptors with statistical significant differences (49). The three groups of regulators were correctly classified in three independent branches. The procedure described here may help identify new chemical compounds with potential uses as plant growth regulators.
Keywords: Chemo-informatics; Plant growth regulators; Molecular descriptors
Introduction
Auxins, cytokinins and gibberellins are by far the most important substances for regulating growth and morphogenesis in plant cell, tissue and organ culture [1-3]. Recently, for instance, in vitro rooting was enhanced in Nicotiana benthamiana by auxin indoleacetic acid [4]. Indolebutyric acid was recommended for shoot and root organogenesis of Eriocephalus africanus, a medicinal and aromatic plant species [5]. Naphthalene acetic acid significantly increased the number of bulblets developed on leaf explants of Scadoxus puniceus [6]. Callus cultures from leaves and young shoots of Taxus globosa were produced with 2,4-dichlorophenoxiacetic acid [7].
Examples of cytokinin use in plant in vitro culture can be found in the following papers. Date palm Phoenix dactylifera L. acclimatization was improved with the use of kinetin [8]. Micropropagation by axillary budding of Quercus ilex was achieved by culturing shoots with zeatin [9]. A reliable protocol was established for in vitro propagation of Artemisia nilagirica with N6-isopenteyladenine [10]. Heuchera villosa petioles were cultured with N6-benzyladenine to induce callus formation [11]. Regarding gibberellins, they have been used to control in vitro morphogenesis of potato [12], bromeliads [13] and peony [14].
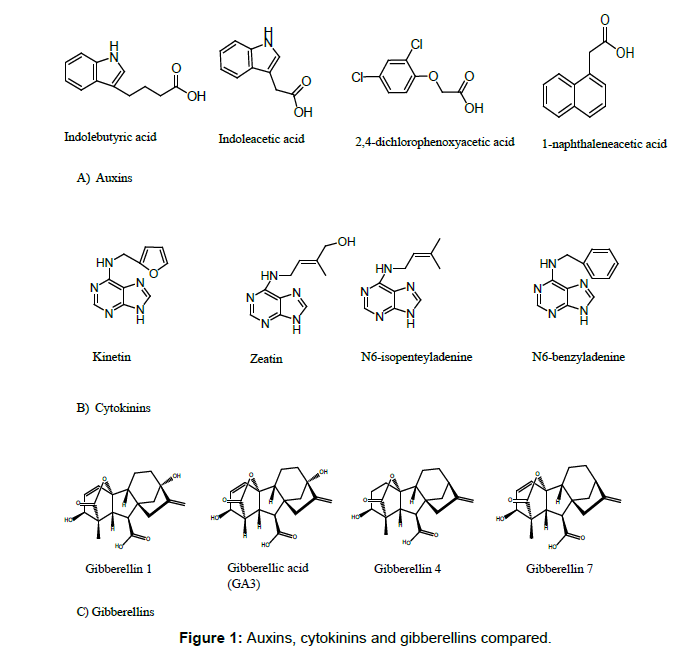
In spite of the physiological effects of auxins, cytokinins and gibberellins have been frequently studied, their chemical dissimilarities to justify their differential impact on plants require more attention. The present study compared the molecular descriptors of indolebutyric acid, indoleacetic acid, 2,4-dichlorophenoxyacetic acid, 1-naphthaleneacetic acid (auxins), kinetin, zeatin, N6-isopentenyl adenine, N6-benzyladenine (cytokinins), GA1, GA3, GA4 and GA7 (gibberellins) (Figure 1).
Materials and Methods
DRAGON software (version 5.5, 2007) and Cambridge Soft Chem Office (version 12, 2010) including ChemDraw and Chem3D were used to calculate 212 molecular descriptors. All data of this study were statistically evaluated using SPSS (Version 8.0 for Windows, SPSS Inc., New York, NY) to perform One - Way ANOVA and Tukey (p=0.05). The overall coefficients of variation (OCV) were calculated as follows: (standard deviation/average) *100. In this formula, we considered the average values of the three growth regulators compared (auxins, cytokinins, gibberellins) to calculate the standard deviation and average. Therefore, the higher the difference between the three groups of chemicals, the higher is the OCV [15]. A hierarchical cluster analysis using the molecular descriptors for auxins, cytokinins and gibberellins was performed. The dendogram was built using average linkage (between groups). Variables were standardized to vary from 0 to 1 according to Kantardzic [16].
Results and Discussion
Even though 49 (out of 212) molecular descriptors showed statistically significant differences among auxins, cytokinins and gibberellins, based on the OCVs in (Table 1), only the numbers of terminal tertiary C (sp3), terminal quaternary C (sp3), ring secondary C (sp3), ring tertiary C (sp3), ring quaternary C (sp3), nitrogen atoms, substituted benzene C (sp2), benzene-like rings, terminal secondary C (sp3), 10-membered rings, and unsubstituted benzene C (sp2) were classified as ¨High¨ OCVs (116.94-173.21%). They indicated a remarkable distinction among these three groups of regulators.
| Auxins | Cytokinins | Gibberellins | Auxins1 | Cytokinins2 | Gibberellins3 | OCV4 | Classification of OCV5 | ||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| IBA | IAA | 2,4-D | NAA | KIN | ZEA | 2IP | BA | GA1 | GA3 | GA4 | GA7 | ||||||
| Number of terminal tertiary C (sp3) | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 5 | 6 | 5 | 5 | 0.00 ± 0.00b | 0.00 ± 0.00b | 5.25 ± 0.25a | 173.21 | High |
| Number of terminal quaternary C (sp3) | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 2 | 1 | 2 | 2 | 0.00 ± 0.00b | 0.00 ± 0.00b | 1.75 ± 0.25a | 173.21 | High |
| Number of ring secondary C (sp3) | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 5 | 5 | 7 | 5 | 0.00 ± 0.00b | 0.00 ± 0.00b | 5.50 ± 0.50a | 173.21 | High |
| Number of ring tertiary C (sp3) | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 5 | 6 | 5 | 5 | 0.00 ± 0.00b | 0.00 ± 0.00b | 5.25 ± 0.25a | 173.21 | High |
| Number of ring quaternary C (sp3) | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 2 | 1 | 2 | 2 | 0.00 ± 0.00b | 0.00 ± 0.00b | 1.75 ± 0.25a | 173.21 | High |
| Number of nitrogen atoms | 1 | 1 | 0 | 0 | 5 | 5 | 5 | 5 | 0 | 0 | 0 | 0 | 0.50 ± 0.29b | 5.00 ± 0.00a | 0.00 ± 0.00b | 150.21 | High |
| Number of substituted benzene C (sp2) | 2 | 2 | 3 | 3 | 0 | 0 | 0 | 1 | 0 | 0 | 0 | 0 | 2.50 ± 0.29a | 0.25 ± 0.25b | 0.00 ± 0.00b | 150.21 | High |
| Number of benzene-like rings | 1 | 1 | 1 | 2 | 0 | 0 | 0 | 1 | 0 | 0 | 0 | 0 | 1.25 ± 0.25a | 0.25 ± 0.25b | 0.00 ± 0.00b | 132.29 | High |
| Number of terminal secondary C (sp3) | 3 | 1 | 0 | 1 | 0 | 0 | 0 | 0 | 5 | 5 | 7 | 5 | 1.25 ± 0.63b | 0.00 ± 0.00b | 5.50 ± 0.50a | 128.14 | High |
| Number of 10-membered rings | 0 | 0 | 0 | 1 | 0 | 0 | 0 | 0 | 1 | 1 | 1 | 1 | 0.25 ± 0.25b | 0.00 ± 0.00b | 1.00 ± 0.00a | 124.9 | High |
| Number of unsubstituted benzene C (sp2) | 4 | 4 | 3 | 7 | 0 | 0 | 0 | 5 | 0 | 0 | 0 | 0 | 4.50 ± 0.87a | 1.25 ± 1.25ab | 0.00 ± 0.00b | 121.19 | High |
| Number of aliphatic secondary C (sp2) | 0 | 0 | 0 | 0 | 0 | 1 | 1 | 0 | 2 | 2 | 0 | 2 | 0.00 ± 0.00b | 0.50 ± 0.29ab | 1.50 ± 0.50a | 114.56 | Medium |
| Number of double bonds | 1 | 1 | 1 | 1 | 0 | 1 | 1 | 0 | 4 | 4 | 3 | 4 | 1.00 ± 0.00b | 0.50 ± 0.29b | 3.75 ± 0.25a | 100 | Medium |
| Number of aliphatic tertiary C (sp2) | 0 | 0 | 0 | 0 | 0 | 1 | 1 | 0 | 1 | 1 | 1 | 1 | 0.00 ± 0.00b | 0.50 ± 0.29ab | 1.00 ± 0.00a | 100 | Medium |
| Number of circuits | 3 | 3 | 1 | 3 | 4 | 3 | 3 | 4 | 15 | 15 | 15 | 15 | 2.50 ± 0.50b | 3.50 ± 0.29b | 15.00 ± 0.00a | 99.23 | Medium |
| Number of oxygen atoms | 2 | 2 | 3 | 2 | 1 | 1 | 0 | 0 | 6 | 6 | 5 | 5 | 2.25 ± 0.25b | 0.50 ± 0.29c | 5.50 ± 0.29a | 92.26 | Medium |
| Number of hydroxyl groups | 1 | 1 | 1 | 1 | 0 | 1 | 0 | 0 | 3 | 3 | 2 | 2 | 1.00 ± 0.00b | 0.25 ± 0.25b | 2.50 ± 0.29a | 91.65 | Medium |
| Number of aromatic bonds | 10 | 10 | 6 | 11 | 15 | 10 | 10 | 16 | 0 | 0 | 0 | 0 | 9.25 ± 1.11a | 12.75 ± 1.60a | 0.00 ± 0.00b | 89.83 | Medium |
| Aromatic ratio | 0.625 | 0.714 | 0.462 | 0.73 | 0.833 | 0.588 | 0.625 | 0.842 | 0 | 0 | 0 | 0 | 0.63 ± 0.06a | 0.72 ± 0.07a | 0.00 ± 0.00b | 87.16 | Medium |
| Number of aromatic C (sp2) | 8 | 8 | 6 | 10 | 9 | 5 | 5 | 11 | 0 | 0 | 0 | 0 | 8.00 ± 0.82a | 7.50 ± 1.50a | 0.00 ± 0.00b | 86.74 | Medium |
| Squared Ghose-Crippen octanol-water partition coefficient. (log^P) | 7.156 | 3.107 | 7.907 | 5.653 | 1.186 | 0.235 | 2.483 | 2.869 | 0.174 | 0 | 2.8 | 2.246 | 5.96 ± 1.06a | 1.69 ± 0.60b | 1.31 ± 0.71b | 86.45 | Medium |
| Number of 5-membered rings | 1 | 1 | 0 | 0 | 2 | 1 | 1 | 1 | 3 | 3 | 3 | 3 | 0.50 ± 0.29b | 1.25 ± 0.25b | 3.00 ± 0.00a | 81.03 | Medium |
| Rotatable bond fraction | 0.138 | 0.087 | 0.158 | 0.08 | 0.111 | 0.133 | 0.103 | 0.1 | 0.02 | 0.021 | 0.019 | 0.02 | 0.12 ± 0.02a | 0.11 ± 0.01a | 0.02 ± 0.00b | 65.65 | Medium |
| Number of 9-membered rings | 1 | 1 | 0 | 0 | 1 | 1 | 1 | 1 | 2 | 2 | 2 | 2 | 0.50 ± 0.29b | 1.00 ± 0.00b | 2.00 ± 0.00a | 65.47 | Medium |
| Number of rings | 2 | 2 | 1 | 2 | 3 | 2 | 2 | 3 | 5 | 5 | 5 | 5 | 1.75 ± 0.25b | 2.50 ± 0.29b | 5.00 ± 0.00a | 55.19 | Low |
| Number of multiple bonds | 11 | 11 | 7 | 12 | 15 | 11 | 11 | 16 | 4 | 4 | 3 | 4 | 10.25 ± 1.11a | 13.25 ± 1.31a | 3.75 ± 0.25b | 53.46 | Low |
| Ghose-Crippen octanol-water partition coefficient (logP) | 2.675 | 1.763 | 2.812 | 2.378 | 1.089 | 0.485 | 1.576 | 1.694 | 0.417 | 0.001 | 1.673 | 1.499 | 2.41 ± 0.23a | 1.21 ± 0.28ab | 0.90 ± 0.41b | 52.92 | Low |
| Number of rotatable bonds | 4 | 2 | 3 | 2 | 3 | 4 | 3 | 3 | 1 | 1 | 1 | 1 | 2.75 ± 0.48a | 3.25 ± 0.25a | 1.00 ± 0.00b | 50.63 | Low |
| Number of hydrogen atoms | 13 | 9 | 6 | 10 | 9 | 13 | 13 | 11 | 22 | 20 | 24 | 22 | 9.50 ± 1.44b | 11.50 ± 0.96b | 22.00 ± 0.82a | 46.84 | Low |
| Number of acceptor atoms for H-bonds (N.O.F) | 2 | 2 | 3 | 2 | 5 | 5 | 4 | 4 | 6 | 6 | 5 | 5 | 2.25 ± 0.25b | 4.50 ± 0.29a | 5.50 ± 0.29a | 40.77 | Low |
| Number of bonds | 29 | 23 | 19 | 25 | 27 | 30 | 29 | 30 | 51 | 48 | 52 | 50 | 24.00 ± 2.08b | 29.00 ± 0.71b | 50.25 ± 0.85a | 40.5 | Low |
| Number of atoms | 28 | 22 | 19 | 24 | 25 | 29 | 28 | 28 | 47 | 44 | 48 | 46 | 23.25 ± 1.89b | 27.50 ± 0.87b | 46.25 ± 0.85a | 37.85 | Low |
| Sum of atomic Sanderson Electronegativities (scaled on Carbon atom) | 28.06 | 22.29 | 20.16 | 24.07 | 25.6 | 29.37 | 28.04 | 28.16 | 47.68 | 44.8 | 48.24 | 46.36 | 23.65 ± 1.67b | 27.79 ± 0.79b | 46.77 ± 0.77a | 37.66 | Low |
| Number of non-H bonds | 16 | 14 | 13 | 15 | 18 | 17 | 16 | 19 | 29 | 28 | 28 | 28 | 14.50 ± 0.65c | 17.50 ± 0.65b | 28.25 ± 0.25a | 36 | Low |
| Number of carbon atoms | 12 | 10 | 8 | 12 | 10 | 10 | 10 | 12 | 19 | 18 | 19 | 19 | 10.50 ± 0.96b | 10.50 ± 0.50b | 18.75 ± 0.25a | 35.95 | Low |
| Sum of Kier-Hall electrotopological states | 36.67 | 33.67 | 38.89 | 35.17 | 35.67 | 37.67 | 32.17 | 36.17 | 65.83 | 63.92 | 58.92 | 59.92 | 36.10 ± 1.11b | 35.4 ± 1.16b | 62.15 ± 1.64a | 34.2 | Low |
| Sum of atomic polarizabilities (scaled on carbon atom) | 18.48 | 14.96 | 14.12 | 16.72 | 17.01 | 18.53 | 18.07 | 19.31 | 30.1 | 28.34 | 30.41 | 29.65 | 16.07 ± 0.97b | 18.23 ± 0.48b | 29.63 ± 0.46a | 34.18 | Low |
| Sum of atomic Van der Waals volumes (scaled on carbon atom) | 17.6 | 14.41 | 13.33 | 16.01 | 16.68 | 17.87 | 17.36 | 18.76 | 28.65 | 27.05 | 28.73 | 28.13 | 15.34 ± 0.93a | 17.67 ± 0.44b | 28.14 ± 0.39b | 33.46 | Low |
| Topological polar surface area using N.O polar contributions | 53.09 | 53.09 | 46.53 | 37.3 | 79.63 | 86.72 | 66.49 | 66.49 | 104.06 | 104.06 | 83.83 | 83.83 | 47.50 ± 3.74b | 74.83 ± 5.03a | 93.95 ± 5.84a | 32.38 | Low |
| Topological polar surface area using N.O.S.P polar contributions | 53.09 | 53.09 | 46.53 | 37.3 | 79.63 | 86.72 | 66.49 | 66.49 | 104.06 | 104.06 | 83.83 | 83.83 | 47.50 ± 3.74b | 74.83 ± 5.03a | 93.95 ± 5.84a | 32.38 | Low |
| Numb of non-N atom | 15 | 13 | 13 | 14 | 16 | 16 | 15 | 17 | 25 | 24 | 24 | 24 | 13.75 ± 0.48c | 16.00 ± 0.41b | 24.25 ± 0.25a | 30.71 | Low |
| Molecular weight | 203.26 | 175.2 | 221.04 | 186.22 | 215.24 | 219.28 | 203.28 | 225.28 | 346.41 | 332.38 | 332.43 | 330.41 | 196.43 ± 10.03b | 215.77 ± 4.65b | 335.41 ± 3.70a | 30.21 | Low |
| Number of 6-membered rings | 1 | 1 | 1 | 2 | 1 | 1 | 1 | 2 | 2 | 2 | 2 | 2 | 1.25 ± 0.25b | 1.25 ± 0.25b | 2.00 ± 0.00a | 28.87 | Low |
| Unsaturation index | 3.585 | 3.585 | 3 | 3.7 | 4 | 3.585 | 3.585 | 4.087 | 2.322 | 2.322 | 2 | 2.322 | 3.47 ± 0.16a | 3.81 ± 0.13a | 2.24 ± 0.08b | 26.03 | Low |
| Ghose-Crippen molar refractivity | 57.654 | 48.452 | 48.366 | 53.816 | 57.686 | 61.575 | 59.8 | 65.295 | 86.414 | 81.913 | 84.078 | 84.887 | 52.07 ± 2.26c | 61.09 ± 1.61b | 84.32 ± 0.94a | 25.28 | Low |
| Sum of conventional bond orders (H-depleted) | 22 | 20 | 17 | 21.5 | 25.5 | 23 | 22 | 27 | 33 | 32 | 31 | 32 | 20.13 ± 1.13c | 24.38 ± 1.14b | 32.00 ± 0.41a | 23.6 | Low |
| Mean electrotopological state | 2.44 | 2.59 | 2.99 | 2.51 | 2.23 | 2.35 | 2.14 | 2.13 | 2.63 | 2.66 | 2.45 | 2.5 | 2.63 ± 0.12a | 2.21 ± 0.05b | 2.56 ± 0.05a | 9.1 | Low |
| Mean atomic van der Waals volume (scaled on carbon atom) | 0.63 | 0.65 | 0.7 | 0.67 | 0.67 | 0.62 | 0.62 | 0.67 | 0.61 | 0.61 | 0.6 | 0.61 | 0.66 ± 0.01a | 0.65 ± 0.01a | 0.61 ± 0.00b | 4.4 | Low |
| Mean atomic polarizability (scaled on carbon atom) | 0.66 | 0.68 | 0.74 | 0.7 | 0.68 | 0.64 | 0.62 | 0.69 | 0.64 | 0.64 | 0.63 | 0.64 | 0.70 ± 0.02a | 0.60 ± 0.02ab | 0.64 ± 0.00b | 4.4 | Low |
2Average information of kinetin, zeatin, N6 – isopentenyladenine and N6-benzyladenine.
3Average information of gibberellin 1, gibberellin 3, gibberellin 4 and gibberellin 7. Results with the same letter are not statistically different (One-Way ANOVA, Tukey, p=0.05).
4Overall coefficient of variation=(Standard deviation/Average)*100. To calculate this coefficient, average values of auxins, cytokinins and gibberellins were considered. The higher the difference among these three averages, the higher the overall coefficient of variation.
5Classification of OCVs: ¨Low¨ from 4.40 to 60.67%; ¨Medium¨ from 60.67 to 116.94% and ¨High¨ from 116.94 to 173.21%.
Table 1: Comparison of molecular descriptors for auxins. cytokinins and gibberellins. IBA: Indolebutyric acid; IAA: Indoleacetic acid; 2,4-D: 2,4-dichlorophenoxyacetic acid; NAA: 1-Naphthaleneacetic acid; KIN: Kinetin; ZEA: Zeatin; 2IP: N6 – isopentenyladenine; BA: N6-benzyladenine; GA1: Gibberellin 1; GA3: Gibberellin 3; GA4: Gibberellin 4; GA7: Gibberellin 7.
Gibberellins contain terminal tertiary C (sp3), terminal quaternary C (sp3), ring secondary C (sp3), ring tertiary C (sp3), and ring quaternary C (sp3) that are not present either in cytokinins or auxins. Gibberellins are also relatively rich in terminal secondary C (sp3) (4.4 times more than auxins =5.50/1.25) and 10-membered rings (4 times more than auxins=1.00/0.25) which are absent in cytokinins.
Cytokinins have 10 times more nitrogen atoms than auxins (5.00/0.50) but this atom is not present in gibberellins. Auxins have 10 times more substituted benzene C (sp2) and 5 times more benzene-like rings than cytokinins (2.50/0.25 and 1.25/0.25, respectively) but these structures are not in gibberellins. Regarding the numbers of unsubstituted benzene C (sp2), auxins average 4.50, cytokinins 1.25 but they are absent in gibberellins.
¨Medium¨ OCVs (60.67 to 116.94%) remarkably distinguished gibberellins from auxins and cytokinins (Table 1). Gibberellins have 3 times more aliphatic secondary C (sp2) and 2 times more aliphatic tertiary C (sp2) than cytokinins (1.50/0.50; 1.00/0.50; respectively) but these types of atoms are not in auxins. Gibberellins are also rich in double bonds (7.5 times more than cytokinins=3.75 /0.50 and 3.75 times more than auxins=3.75/1.00), circuits (4.3 times more than cytokinins=15.0/3.5 and 6 times more than auxins=15.0/2.5), and oxygen atoms (11 times more than cytokinins=5.50/0.50 and 2.4 times more than auxins=5.50/2.25).
The numbers of hydroxyl groups, 5-membered and 9-membered rings are also higher in gibberellins: 10 times more hydroxyl groups than cytokinins (2.50/0.25) and 2.5 times more than auxins (2.50/1.00); 2.4 times more 5-membered rings than cytokinins (3.00/1.25) and 6 times more than auxins (3.00/0.50); and 2 times more 9-membered rings than cytokinins (2.00/1.00) and 4 times more than auxins (2.00/0.50). Contrastingly, gibberellins do not have either aromatic bonds (cytokinins: 12.75; auxins: 9.25) or aromatic C (sp2) (cytokinins: 7.50; auxins: 8.00). Aromatic ratio is cero in gibberellins while 0.72 in cytokinins and 0.63 in auxins. On the other hand, the rotatable bond fraction is lower in gibberellins (0.02) compared to auxins (0.12) or cytokinins (0.11). It is important to note the squared Ghose-Crippen octanol-water partition coefficient is remarkably higher in auxins (5.96) in comparison with cytokinins (1.69) and gibberellins (1.31).
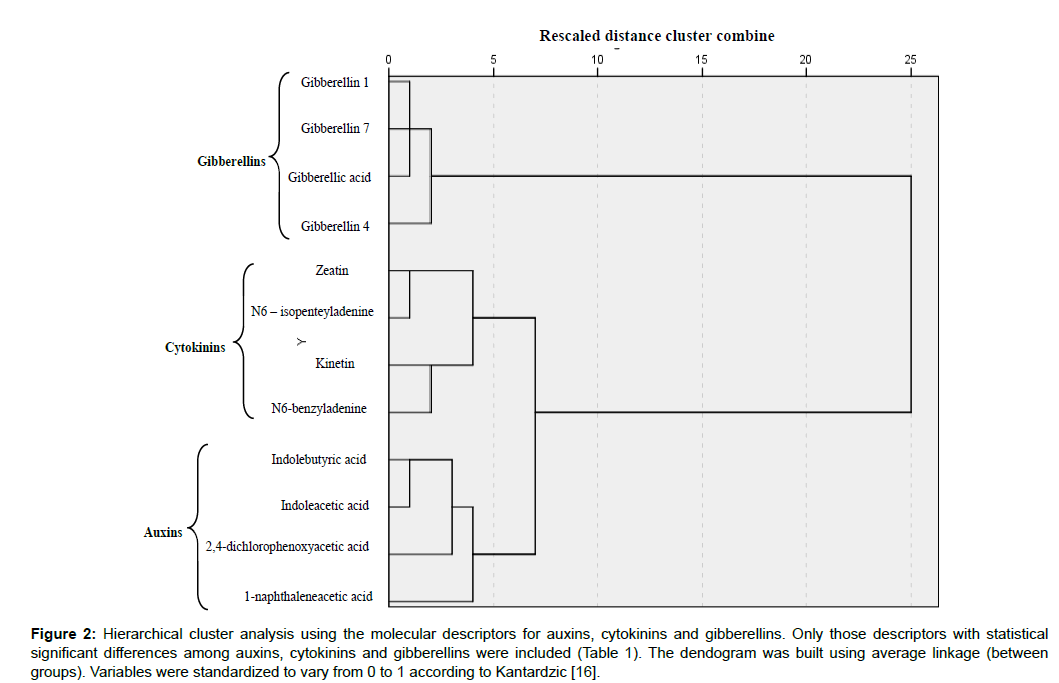
Descriptors shown in Table 1 were used to generate the dendogram shown in Figure 2 which correctly classified the three groups of regulators in three independent branches. Molecular descriptors have been applied to describe biological activities, in many studies showing their applicability as an attractive tool for efficient (e.g.) drug design process [17-19].
Figure 2: Hierarchical cluster analysis using the molecular descriptors for auxins, cytokinins and gibberellins. Only those descriptors with statistical significant differences among auxins, cytokinins and gibberellins were included (Table 1). The dendogram was built using average linkage (between groups). Variables were standardized to vary from 0 to 1 according to Kantardzic [16].
To end we would like to emphasize the effectiveness of the chemoinformatics procedure described here to differentiate auxins, cytokinins and gibberellins and also in the search for new plant growth regulators with potential applications in modern in vitro culture and agriculture. Molecular descriptors of new chemical compounds can be determined and included in the dendogram shown in Figure 2. If new chemicals are located, for instance, near auxins they can be regarded as potential auxinlike compounds, although this should be later tested experimentally.
Author Contribution
I.A., D.G., L.P. and J.C.L. designed the research, analyzed the data and wrote the paper. J.C.L. had primary responsibility for the final content. All authors have read and approved the final manuscript.
Acknowledgements
This research was supported by the Bioplant Centre (University of Ciego de Ávila, Cuba).
References
- Salisbury FB, Ross CW (1992) Plant Physiology (In Spanish). (4th Edtn), Wadsworth Publishing, California.
- Machakova I, Zazimalova E, George EF (2008) Plant growth regulators I: Introduction; auxins, their analogues and inhibitors, In: George EF, Hall M, De Kerk GJ (eds) Plant Propagation by Tissue Culture. (3rd Edtn), Springer, Dordrecht 175-204.
- van Staden J, Zazimalova E, George EF (2008) Plant growth regulators II: cytokinins, their analogues and antagonists. In: George EF, Hall M, De Kerk GJ (eds) Plant Propagation by Tissue Culture. (3rd Edtn), Springer, Dordrecht 205-226.
- Esserti S, Faize M, Rifai LA, Smaili A, Belfaiza M, et al. (2017) Media derived from brown seaweeds Cystoseira myriophylloides and Fucus spiralis for in vitro plant tissue culture. Plant Cell Tiss Org Cult 128: 437-446.
- Madzikane-Mlungwana O, Moyo M, Aremu AO, Plíhalová L, Doležal K, et al. (2017) Differential responses to isoprenoid, N 6-substituted aromatic cytokinins and indole-3-butyric acid in direct plant regeneration of Eriocephalus africanus. Plant Grow Reg 82: 103-110.
- Naidoo G (2010) Ecophysiological differences between fringe and dwarf Avicennia marina mangroves. Trees 24: 667-673.
- Tapia N, Zamilpa A, Bonfill M, Ventura E, Cruz-Vega D, et al. (2013) Effect of the culture medium and biotic stimulation on taxane production in Taxus globosa Schltdl in vitro cultures. Acta Physiol Plant 35: 3447-3455.
- Hassan MM (2017) Improvement of in vitro date palm plantlet acclimatization rate with kinetin and Hoagland solution. In: Al-Khayri JM, Jain SM, Johnson DV (eds) Date Palm Biotechnology Protocols Volume I: Tissue Culture Applications. Springer New York, New York 185-200.
- Martínez MT, Corredoira E, Vieitez AM, Cernadas MJ, Montenegro R, et al. (2017) Micropropagation of mature Quercus ilex L. trees by axillary budding. Plant Cell Tiss Org Cult 131: 499-512.
- Shinde S, Sebastian JK, Jain JR, Hanamanthagouda MS, Murthy HN (2016) Efficient in vitro propagation of Artemisia nilagirica var. nilagirica (Indian wormwood) and assessment of genetic fidelity of micropropagated plants. Physiol Mol Biol Plants 22: 595-603.
- Zhao HQ, He QH, Song LL, Hou MF, Zhang ZG (2017b) In Vitro culture of Heuchera villosa ‘Caramel’. HortScience 52: 622-624.
- Ali S, Khan N, Nouroz F, Erum S, Nasim W, et al. (2018) In vitro effects of GA3 on morphogenesis of CIP potato explants and acclimatization of plantlets in field. In Vitro Cell Devel Biol-Plant 54: 104-111.
- dos Santos DS, Cardoso-Gustavson P, Nievola CC (2017) Stem elongation of ornamental bromeliad in tissue culture depends on the temperature even in the presence of gibberellic acid. Acta Physiol Plant 39: 230.
- Zhao D, Xue Y, Shi M, Tao J (2017a) Rescue and in vitro culture of herbaceous peony immature embryos by organogenesis. Sci Hort 217: 123-129.
- Lorenzo JC, Yabor L, Medina N, Quintana N, Wells V (2015) Coefficient of variation can identify the most important effects of experimental treatments. Not Bot Horti Agrobo Cluj-Nap 43: 287-291.
- Kantardzic M (2003) Data Mining: Concepts, Models, Methods and Algorithms. (1st Edtn) Wiley, New Jersey.
- Casanola-Martin GM, Khan MTH, Marrero-Ponce Y, Ather A, Sultan S, et al. (2007) TOMOCOMD-CARDD descriptors-based virtual screening of tyrosinase inhibitors. 1. Evaluation of different classification model combinations using group, atom-type and total bilinear indices. Bioorg Med Chem 15: 1483-1503.
- Santana KD, PhamThe H, Borroto OMR, Puris A, Thu HLT, et al. (2017) A Two QSAR way for antidiabetic agents targeting using α-amylase and α-glucosidase inhibitors: model parameters settings in artificial intelligence techniques. Lett Drug Des Disc 14: 862-868.
- The HP, Martin GMC, Santana KD, Hai NN, Ngoc NT, et al. (2017) Quantitative structure-activity relationship analysis and virtual screening studies for identifying HDAC2 inhibitors from known HDAC bioactive chemical libraries. SAR QSAR Environ Res 28: 199-220.
 Spanish
Spanish  Chinese
Chinese  Russian
Russian  German
German  French
French  Japanese
Japanese  Portuguese
Portuguese  Hindi
Hindi