Research Article, Clin Dermatol Res J Vol: 2 Issue: 2
Gene Expression Profile Analysis and Identification of the Effects Triggered by Essential Sandalwood Oil on Human Skin Explants
Paul Moretta1, Rémi Cléro1, Philippe Benech2 and Thomas Bader2*
1SANTANOL, Couslon Way, Canning Vale, WA, Australia
2GENEX, Chemin de Saulxier, Longjumeau, France
*Corresponding Author : Thomas Bader
GENEX, Chemin de Saulxier, Longjumeau, France
Tel: +33164477915
E-mail: th.bader@laboratoire-genex.fr
Received: September 28, 2017 Accepted: November 17, 2017 Published: November 22, 2017
Citation: Moretta P, Cléro R, Benech P, Bader T (2017) Gene Expression Profile Analysis and Identification of the Effects Triggered by Essential Sandalwood Oil on Human Skin Explants. Clin Dermatol Res J 2:2. doi: 10.4172/2576-1439.1000120
Abstract
Sandalwood oil is an essential oil obtained from the tree wood of various species of Santalum spp by steam distillation, and has a history of medicinal use besides as a fragrance. However the
mechanisms involved in its activity are still unclear. In the present report, Sandalwood oil (S. album) was applied to living human skin explants to evaluate the sustained activity of the oil through
gene expression profiling. Transcriptomic analysis showed a number of metabolic pathways and biological activities triggered by treatments previously described for aloe, curcumin and luteolin. No inflammatory responses accompanied the beneficial activities. The most intriguing impact was the natural effect on the retinoic acid (vitamin A) metabolism.
Furthermore, we found that mechanisms normally acting in more specialized cell types such as immune cells, adipocytes, nerve cells, or hair follicles were activated in human skin explants in
response to sandalwood oil. Thus, the oil may have the potential of further beneficial activities such as its antibacterial activity among other protective mechanisms.
These results corroborate at a molecular level the rationale for clinical studies with sandalwood oil and support recent studies of patients with eczema (atopic dermatitis) or acne, which is often
accompanied by bacterial growth on skin of Propionibacterium acnes.
Keywords: Sandalwood oil; Skin; Gene activation; Microarray analysis; Expression profiling; Transcriptome; Plant extracts
Introduction
Sandalwood oil (SO) is an essential oil obtained from the wood of various species of Santalum spp. Since ancient times this essential oil served for religious purposes as an incense and used in fragrances, flavorings, and as a traditional medicine for use as an antidepressant, anti-inflammatory, anti-fungal, astringent, sedative, as an insecticide, and antiseptic [1]. SO is a mixture of compounds with similar structural and physical characteristics but contains primarily a sesquiterpene alcohol molecule called santalol, which has two isomeric forms, α-santalol (41-55%) and β-santalol (16-24%), both of which are volatile compounds with a unique aroma and are the primary active components. α-santalol has proved to be a potential chemopreventive agent against UVB-induced skin tumor development in mice [2], while β-santalol was investigated for its antiviral activity against influenza A/HK (H3N2) virus in MDCK cells [3]. Additionally, purified α-santalol and β-santalol were able to suppress secretion of key cytokine-related factors and proinflammatory arachidonic acid metabolite production in skin cells. Lipopolysaccharides stimulated the release of 26 cytokines and chemokines, 20 of which were substantially suppressed by simultaneous exposure to either of the two sandalwood compounds and to ibuprofen [4]. In a study on a cohort of radiotherapy patients, turmeric- and sandal oil-based cream was effective in preventing radiation-induced dermatitis. The damage is mainly mediated via indirect effects, where the generation of free radicals, resulting from the radiolysis of water, causes damage to macromolecules such as DNA, proteins and lipids [5].
SO showed insecticidal activity with LD50 values (μg/fly) of 2.18 and of 5.61 against male and female drosophila flies, respectively [6]. In vivo analysis using a rodent model confirmed the anti-plasmodial potential of subcutaneously administered sandalwood oil [7]. It also is an effective repellent against spider mites [8].
Other minor molecules are derivatives such as the aldehydes santalal and cyclo-santalal, and traces of santalene, curcumene and bergamotene. The latter two terpenes can also be found in essential oils from different plants. For instance, curcuminoids from the curcuma root or sage possess anti-oxidant, anti-inflammatory and anti-mutagenic properties and protect the body from mutagens such as smoke and other pollutants [9]. Bergamotene itself is found in a variety of plants, from basil [10], and the Brazilian tree Pau Santo (Kielmeyera coriacea) to orchids [11], and has been used for its antimicrobial activity [12].
Presently, seven clinical trials are under way which utilize SO in treatments of common warts, the infection molluscum contagiosum in children, caused by a poxvirus, atopic dermatitis (eczema) and acne, and oral mucositis or plaque psoriasis (clinicaltrials.org). In preliminary results treating eczema with SO, 68% of patients achieved an IGA score of “much improved” or “very much improved” with a minimum 2-grade improvement after eight weeks of treatment (NCT02871479).
The purpose of the present study was to establish at the molecular level, a genetic profile of the biological activity of essential sandalwood oil from Santalum album, produced under standardized conditions. The model system under investigation is based on living human skin explants. This protocol permits to compare ex vivo the effects of SO at the histological level as well as the changes of gene expression as a function of SO concentration and duration of treatment. Using proprietary dedicated software, PredictSeach®, functional correlations between genes were identified allowing the generation of biological networks related to SO activities.
Materials and Methods
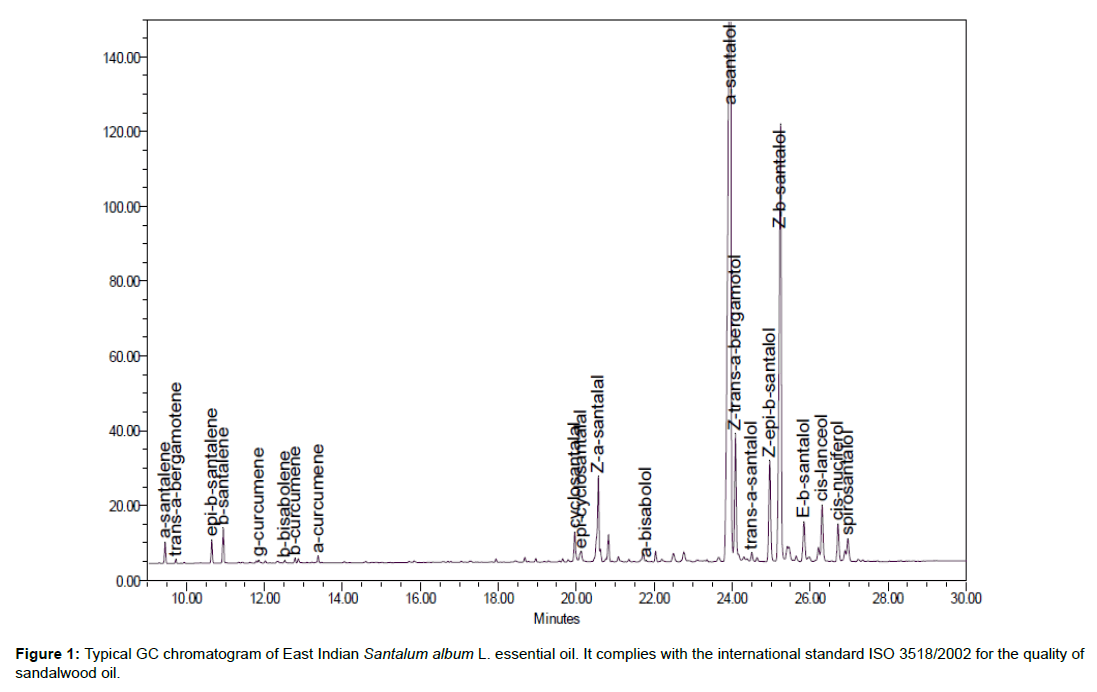
The wood was obtained from managed plantations of Santalum album grown in the Kununurra district of Western Australia. The plantations were owned by Santanol. The essential oil was extracted from the rich heartwood of plantation trees that were 15 years old. Oil was obtained through the steam distillation of wood chips loaded into commercial size stainless steel distillation pots. The effluent was cooled and condensed into a collection vessel and the top layer containing the oil was collected. The quality of the oil complies with that set out in ISO 3518/2002 – Oils of Sandalwood (Santalum album L.). A typical chromatogram is shown in Figure 1, and the extracted SO presented a composition as shown in Table 1.
| Name | % |
|---|---|
| Z-a-santalol | 46.35 |
| Z-b-santalol | 19.11 |
| Z-trans-a-bergamotol | 5.66 |
| Z-epi-b-santalol | 3.91 |
| Z-a-santalal | 3.86 |
| cis-lanceol | 2.8 |
| cis-nuciferol | 1.78 |
| E-b-santalol | 1.63 |
| Spirosantalol | 0.98 |
| Cyclosantalal | 0.93 |
| b-santalene | 0.84 |
| epi-cyclosantalal | 0.58 |
| epi-b-santalene | 0.54 |
| a-santalene | 0.51 |
| trans-a-santalol | 0.33 |
| a-bisabolol | 0.18 |
| a-curcumene | 0.17 |
| trans-a-bergamotene | 0.1 |
Table 1: Typical composition of essential oils after the standardized extraction of sandalwood.
The study was approved by an ethics committee and performed in accordance with the Declaration of Helsinki after the patient had given informed consent. A full-thickness human skin biopsy was obtained after plastic surgery from a healthy female donor (49-yearold Caucasian). The abdoplasty was immediately transported at room temperature in minimum essential medium 1X (MEM, Gibco life technologies™; Paisley, UK) to BIO-EC laboratory. The hypodermis was removed from the skin, and circular explants (~1cm diameter, 0.2cm thickness, 180mg) were then excised using a sample punch. Explants, dermis face down, were directly placed in a liquid– air interface in BIO-EC’s Explant Medium (BEM), containing gentamicin and fungizone for 24h before treatment. Skin samples were maintained under typical cell culture conditions (37°C in 5% CO2), and half of the medium (1ml) was refreshed every other day.
Samples were taken for each timepoint. Triplicates were removed at 9h and 24h for genomic expression analysis and histology, and on day 8 for morphological evaluation. Treatment consisted of excipient alone (sweet almond oil) as control, or 0.1, 0.2, 0.4% (P3), 1.0 and 2% (P1) Santalum album oil in excipient, on day 0 (D0), day 1, day 4 and day 6 on the basis of 2mg per explant.
Samples were cut in half for storage in RNAlater (Qiagen) or fixation in buffered formol. For histology, samples were dehydrated and impregnated in paraffin, and embedded according to the SOP H-153 using a Leica EG 1160 embedding station. 5-μm-thick sections were made according to the SOP H-173 with a Leica RM 2125 Minottype microtome, and the sections were then mounted on Superfrost® Plus silanized glass slides. Microscopical observations were realized using a Leica DMLB or Olympus BX43 microscope. Pictures were digitized with a numeric DP72 Olympus camera with Cell D storing software. General morphology was assessed on controls D-1, D0, and D8, and treated samples on D8 after staining of paraffinized sections according to Masson’s trichrome, Goldner variant.
Total RNA was extracted with Relia Prep Tissue Mini prep system (fibrous) from Pro mega. RNA (40ng) of each sample was used for reverse transcription, amplification and Cy3 labeling, using the Low Input Labeling kit, one-color (Agilent Technologies). All cRNAs were hybridized to human whole genome microarrays (Agilent Technologies). Microarray data were quantified and normalized according to Agilent protocol (GeneExtraction Feature V10.5.5.1), and deposited in the public domain (GEO Submission GSE106734). Subsequently, the fold-change (FC) between treated versus untreated condition was calculated by the following ratios: transcript intensity of treated samples versus transcript intensity of control samples. The selection of genes of interest were identified in sequential steps: a) Controls were considered unchanged if their expression was between 0.8 and 1.25 fold-change at two time points, 9h (early) and 24h (late). b) In order to take into account the effect of the tested product (P), only gene transcripts that were not subjected to modulation caused by the excipient (almond oil) were considered. Therefore, genes with an FC ≥ 1.25 between excipient and control values (intensities higher than background) in at least two out of three similar conditions (for each permutation between treated samples and controls), were removed.
The remaining genes were then filtered for either FC ≥ 1.25 and <1.45, and for FC ≥ 1.45 between P and control values (intensities higher than background) in at least two out of three similar conditions (for each permutation between all treated samples and controls) for either P1 (2%) or P3 (0.4%) at either one (9h or 24h) or both time points. P1 and P3 were chosen based on the histology results.
Selected genes were subjected to functional analysis by GENEX’ proprietary PredictSearch® program in order to identify the induced biological effects. PredictSearch® is a powerful text mining software application that identifies correlations between genes and biological processes/diseases within all scientific publications citied in the PubMed database.
Results and Discussion
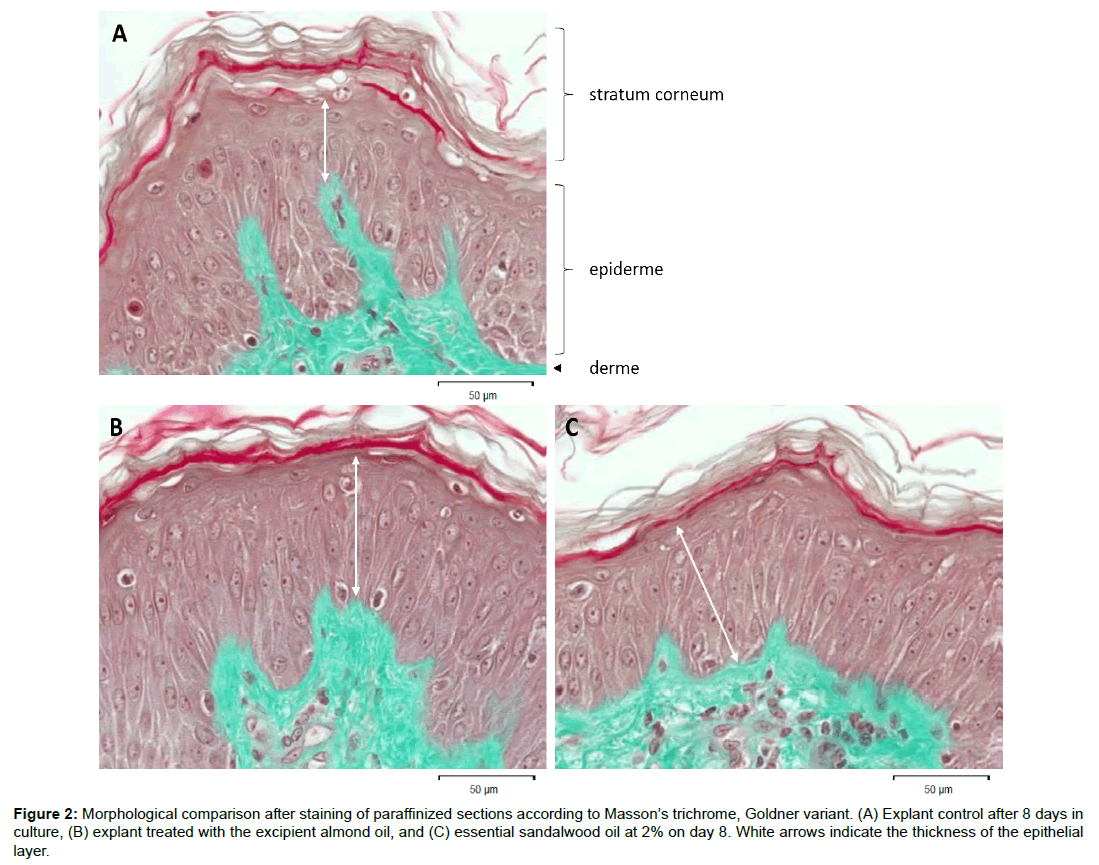
The selection resulted in 65 genes, which were activated as a consequence of sandalwood oil treatment. The complete absence of any inflammatory genes and few immune response-related genes indicated the oil was very well tolerated. It is important to note that for specific plant species, particular characteristics exist in their content of secondary metabolites (phenolics, alkaloids, steroids, terpenoids, acetophenones) according to the geographical location between families. Of the different Santalum species, the SO under investigation comes from Santalum album, which contains no farnesol, compared to the native Australian Santalum spicatum that may contain 5-30% of this contact allergen [13]. Figure 2 compares the morphology of explants on D8 between controls or treated either with excipient or 2% essential oil. There are no discernible alterations but an increase of the epithermal layer following treatment.
Figure 2: Morphological comparison after staining of paraffinized sections according to Masson’s trichrome, Goldner variant. (A) Explant control after 8 days in culture, (B) explant treated with the excipient almond oil, and (C) essential sandalwood oil at 2% on day 8. White arrows indicate the thickness of the epithelial layer.
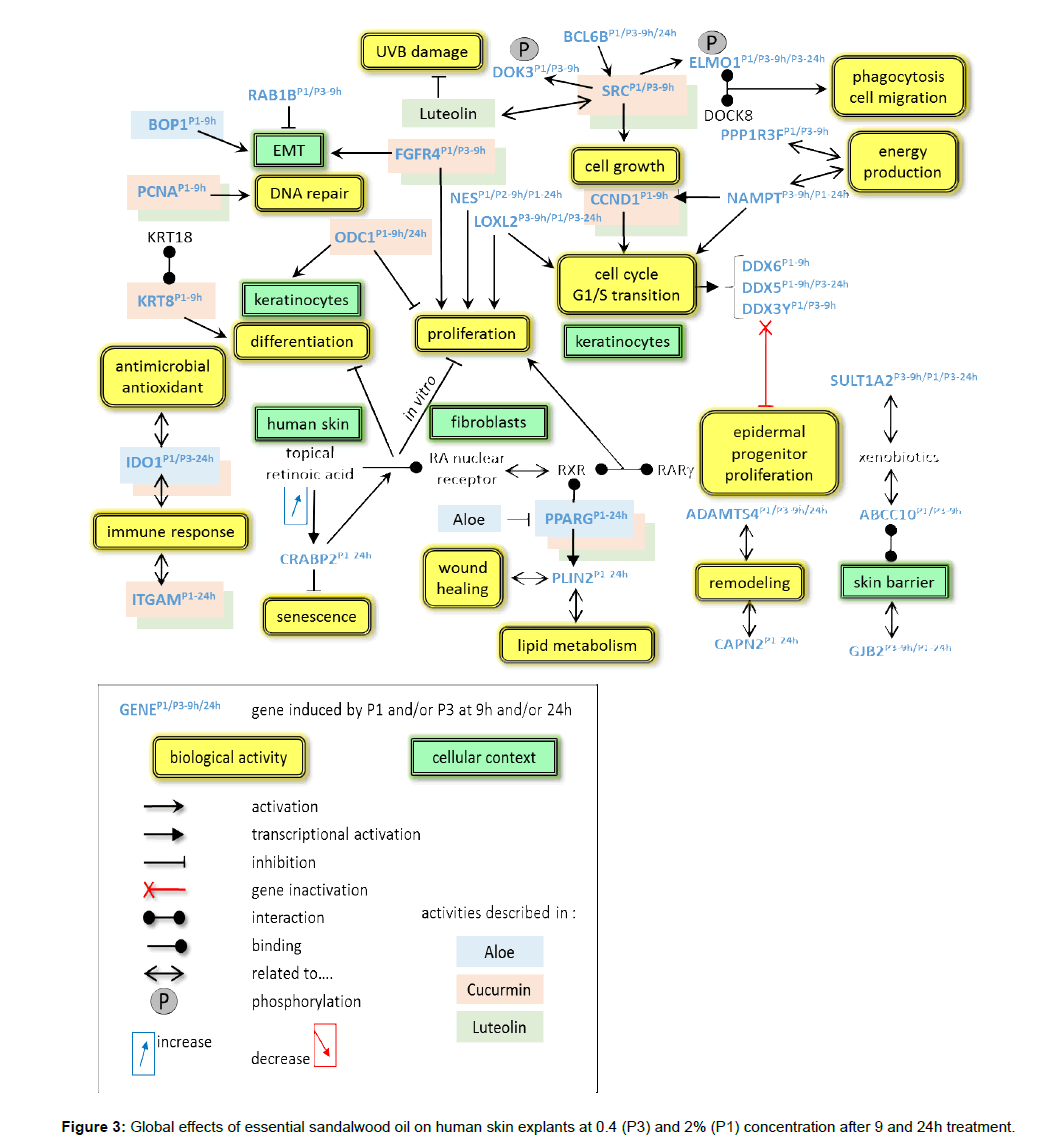
The global overview shows that a retinoic acid (RA) and survival pathway, differentiation and proliferation are the predominant motives as 40 genes related to these processes, partially overlapping, are induced. Interestingly, several activities known from other essential oils or plant extracts can be found in this study, including retinoic acid-related pathways with activities similar to medicinal drugs under development. Table 2 lists the most important genes and the induced fold-changes in this study. The overall activities are summarized in Figure 3 and described below.
| Gene Name | P1 (2%) 9h | SD | P3 (0.4%) 9h | SD | P1 (2%) 24h | SD | P3 (0.4%) 24h | SD |
|---|---|---|---|---|---|---|---|---|
| ABCC10 | 1.85 | 0.40 | 1.58 | 0.13 | 1.07 | 0.19 | 0.84 | 0.09 |
| ADAMTS4 | 1.29 | 0.25 | 1.61 | 0.47 | 1.50 | 0.19 | 1.37 | 0.13 |
| BCL6B | 1.49 | 0.49 | 1.14 | 0.32 | 1.43 | 0.12 | 1.55 | 0.13 |
| BOP1 | 1.40 | 0.06 | 1.07 | 0.08 | 1.05 | 0.12 | 1.01 | 0.08 |
| CAPN2 | 1.20 | 0.23 | 1.08 | 0.22 | 1.34 | 0.06 | 1.10 | 0.15 |
| CCND1 | 1.34 | 0.12 | 0.99 | 0.15 | 0.94 | 0.14 | 0.93 | 0.10 |
| CRABP2 | 0.93 | 0.27 | 0.83 | 0.15 | 1.49 | 0.17 | 1.09 | 0.15 |
| DDX3Y | 1.56 | 0.11 | 1.37 | 0.16 | 1.13 | 0.13 | 0.90 | 0.15 |
| DDX5 | 1.37 | 0.14 | 1.14 | 0.16 | 1.14 | 0.12 | 1.29 | 0.14 |
| DDX6 | 1.29 | 0.06 | 1.10 | 0.15 | 1.02 | 0.13 | 0.91 | 0.12 |
| DOK3 | 1.58 | 0.20 | 2.62 | 0.72 | 1.03 | 0.27 | 0.91 | 0.17 |
| ELMO1 | 1.48 | 0.31 | 1.81 | 0.58 | 0.88 | 0.23 | 1.37 | 0.18 |
| FGFR4 | 1.42 | 0.16 | 1.79 | 0.40 | 0.96 | 0.16 | 1.03 | 0.06 |
| GJB2 | 0.91 | 0.49 | 1.57 | 0.15 | 1.40 | 0.18 | 0.93 | 0.28 |
| IDO1 | 0.87 | 0.19 | 0.85 | 0.17 | 1.33 | 0.12 | 1.35 | 0.16 |
| ITGAM | 0.94 | 0.34 | 0.93 | 0.25 | 1.38 | 0.11 | 1.04 | 0.11 |
| KRT8 | 1.49 | 0.22 | 1.08 | 0.18 | 0.91 | 0.18 | 0.83 | 0.17 |
| LOXL2 | 1.01 | 0.10 | 1.41 | 0.30 | 1.58 | 0.24 | 1.43 | 0.17 |
| NAMPT | 0.93 | 0.11 | 1.35 | 0.09 | 1.32 | 0.06 | 1.12 | 0.15 |
| NES | 1.64 | 0.61 | 1.29 | 0.48 | 1.37 | 0.11 | 1.04 | 0.30 |
| ODC1 | 1.39 | 0.03 | 1.01 | 0.25 | 1.45 | 0.32 | 1.07 | 0.31 |
| PCNA | 1.46 | 0.18 | 1.01 | 0.13 | 1.02 | 0.12 | 0.91 | 0.10 |
| PLIN2 | 0.83 | 0.17 | 0.89 | 0.11 | 1.35 | 0.12 | 1.11 | 0.16 |
| PPARG | 1.18 | 0.16 | 0.99 | 0.29 | 1.57 | 0.19 | 1.20 | 0.31 |
| PPP1R3F | 1.42 | 0.11 | 1.65 | 0.32 | 0.98 | 0.21 | 1.06 | 0.27 |
| RAB1B | 2.32 | 0.03 | 1.74 | 0.35 | 0.97 | 0.10 | 0.82 | 0.05 |
| SRC | 1.38 | 0.12 | 1.35 | 0.10 | 0.96 | 0.13 | 0.96 | 0.15 |
| SULT1A2 | 1.00 | 0.17 | 1.33 | 0.26 | 1.34 | 0.13 | 1.41 | 0.14 |
Table 2: Mean fold-change induced by sandalwood oil in at least two out of three explants. Two concentrations were tested and results of 9h and 24h treatment are shown. Significant changes of expression are in bold.
The following distinct pathways could be attributed to the induced genes: a RA signaling pathway, mechanisms concerning cell cycle control, proliferation and differentiation, including DNA repair, remodeling activities with cell structure changes, and protective activities.
Several signature genes of the RA pathway are induced after 24h with 2% SO: CRABP2 (cellular retinoic acid binding protein 2), PLIN2 (perilipin 2), and PPARG (peroxisome proliferator activated receptor gamma). The protein Crabp2 is a cytosol-to-nuclear shuttling protein. It is involved in the retinoid signaling pathway, and the classical function of Crabp2 is to directly deliver RA to its nuclear receptor (RAR), which in turn induces the expression of multiple anti proliferative genes [14]. Consequently, Crabp2 might potentially act as a negative regulator that limits cellular senescence in fibroblasts [15,16]. PPARs are a family of nuclear receptors, which can heterodimerize with retinoid X receptors to regulate transcription of various genes, including PLIN2 in adipocyte differentiation and lipid metabolism. The protein is the major cAMP-dependent protein kinase substrate in adipocytes and, when unphosphorylated, may play a role in the inhibition of lipolysis [17]. It is also a target gene of PPARG which binds directly to the PPRE site on the promoter [18].
When topically applied, RA induces keratinocyte proliferation and upregulates CRABP2, a signal transduced by RXR alpha/RAR gamma heterodimers in suprabasal keratinocytes, which, in turn, stimulate proliferation of basal keratinocytes [19,20]. The analysis of stimulated gene expression suggests that SO treatment has a significant effect on cell growth and the differentiation processes. The signaling events that correspond to the retinoic acid pathway (CRABP2, PPARG, perilipin 2) decrease cell growth and promote differentiation. Furthermore, the drug Troglitazone®, one of the thiazolidinedione (TZD) class of anti-diabetic drugs and a ligand for PPARG, inhibits CCND1 (cyclin D1) expression. Troglitazone® and other PPARG ligands have been shown to inhibit cell proliferation and induce cell cycle arrest in a variety of cancer cells as well as to inhibit mitogen-induced cellular proliferation in normal mouse skin primary keratinocytes [21].
Most interesting are activities similar to that with the drug Rambazole (Talarozole, Barrier Therapeutics, Geel, Belgium) currently under development for its interference in retinoic acid (vitamin A)-related pathways. Rambazole increases intracellular levels of endogenous all-trans RA through inhibition of cytochrome P450-dependent all-trans-retinoic acid catabolism. It is in clinical development for the treatment of psoriasis and acne [22]. Wellknown effects of RA are normalization of aberrant epithelial growth and differentiation. Hence, Rambazole may be utilized in the treatment of plaque psoriasis [23]. Other skin diseases such as acne or keratinization disorders might benefit from this activity.
More genes, which code for proteins regulating proliferation and differentiation, are detectable: ODC1 (ornithine decarboxylase 1), RAB1B (member RAS oncogene family) and BCL6B (B-cell CLL/lymphoma 6B), all have been shown to have tumor suppressor activity in various cancer models [24-26]. NAMPT (nicotinamide phosphoribosyltransferase, visfatin) belongs to the nicotinic acid phospho-ribosyltransferase (NAPRTase) family and is thought to be involved in many important biological processes, including metabolism, stress response and anti-aging by activating sirtuin 1 [27]. NAMPT is a rate-limiting enzyme in the NAD(+) salvage pathway of NAD(+), which serves as a transfer molecule for electrons, thereby acting as a key cofactor for energy production, an activity shared with PPP1R3 (protein phosphatase 1 regulatory subunit 3F) expressing a glycogen-binding protein [28]. NAMPT activates G1-S phase cell cycle progression by upregulation of CCND1 [29]. DDX5 (DEAD-box helicase) is essential for normal cell proliferation, and the transition from G1 to S/G2 phase is accompanied by an increase of DDX5 protein concentration in the [30]. Two more members of the same family, DDX3Y and DDX6, were induced. The former is found in P-bodies and stress granules, and functions in translation suppression and mRNA degradation. It is required for microRNA-induced gene silencing. The latter, DDX6 is necessary for maintaining adult progenitor cell function and its loss results in premature differentiation and decreased proliferation of epidermal progenitor cells [31].
FGFR4 (fibroblast growth factor receptor 4) encodes a member of the fibroblast growth factor receptor family, where the amino acid sequence is highly conserved between members and throughout evolution. Functional assays have demonstrated that FGFR4 can induce proliferation, invasion, and epithelial-mesenchymal transition (EMT) after FGF stimulation. Indeed, FGFR4 inhibitors can suppress proliferation, invasion and induce apoptosis [32]. LOXL2 (lysyl oxidase like 2) belongs to a gene family, where the prototypic member is essential for the biogenesis of connective tissue. The effect of LOXL2 on cell cycle and apoptosis-related components was has been demonstrated through the silencing of LOXL2 expression, which activates the FAK/Akt/mTOR signaling pathway [33].
BOP1 (block of proliferation 1) has an important role in hepatocellular carcinoma invasiveness and metastasis potentials through inducing epithelial to mesenchymal transition, and promoting actin cytoskeleton remodeling. BOP1 is a novel nucleolar protein involved in rRNA processing and ribosome assembly. Kinase activities of the G(1)-specific CDK2 and CDK4 complexes were downregulated in cells expressing BOP1DELTA, whereas levels of the CDK inhibitors p21 (CDKN1A cyclin-dependent kinase inhibitor 1A) and p27 (CDKN1B cyclin-dependent kinase inhibitor 1B) were concomitantly increased [34]. The conserved protein Bop1 is essential for viability. Lower transcript levels result in defects in rRNA processing and developmental abnormalities that are consistent with its predicted role in ribosome biogenesis [35].
The proliferating cell nuclear antigen (PCNA) is found in the nucleus and is a cofactor of DNA polymerase delta. It acts as a homotrimer and helps increase the processivity of leading strand synthesis during DNA replication, however in response to DNA damage, this protein is ubiquitinated and is involved in the RAD6-dependent DNA repair pathway [36]. NES (nestin) is a key regulator of various extracellular proteins that play important roles in cell growth and differentiation. Overexpression promotes the embryonic development of heart and brain through the regulation of cell proliferation [37]. Nestin is found only in intramesenchymal skin, and stimulation of organ-cultured human scalp skin with the adipokine leptin increased the number of nestin+ cells in these intramesenchymal skin locations [38]. The gene SRC (protooncogene, non-receptor tyrosine kinase Src) may play a role in the regulation of embryonic development and cell growth and its activity can be inhibited by phosphorylation through c-SRC kinase.
According to the kinetics in this study, it is possible that these effects are initiated sequentially. For instance, within the skin granular layer, proliferation of undifferentiated cells is followed by DNA repair activity (to preserve the integrity of growing cells), and growth arrest (inducing some apoptosis-related genes), which in turn allows cells to differentiate. These activities concern cellular structures such as those belonging to the cytoskeleton, and might impact the extra cellular matrix. This notion is supported by activated genes that are involved in cellular architecture and remodeling with the help of proteins like keratins and proteases.
Cellular structural changes and remodeling activities are reflected in the increase of genes like KRT8. The protein keratin 8 plays a role in maintaining cellular structural integrity and also functions in signal transduction and cellular differentiation. ELMO1 (engulfment and cell motility 1) encodes a member of the engulfment and cell motility protein family. These proteins interact with DOCK8 (dedicator of cytokinesis) protein to promote phagocytosis and cell migration. ELMO1 is also a target of phosphorylation by Src [39]. ADAMTS4 (a disintegrin and metallopeptidase with thrombospondin type 1 motif 4) is responsible for the degradation of specific proteoglycans [40], while calpain 2 (CAPN2) belongs to calcium-activated, nonlysosomal, intracellular cysteine proteases. Calpain activity is implicated in important cellular processes including cytoskeletal remodeling, apoptosis and survival [41]. Connexins such as GJP2 (gap junction protein 2) are involved in cell motility and wound healing and GJP2 enhances cell motility [42].
Another facet is protective measures triggered at the molecular level, which target a host of conditions from inflammation to autoimmune disorders, infections and other defense systems.
Induced protective activities can be found in several modulated genes: DOK3 (docking protein 3), a negative regulator of tolllike receptor signaling by limiting LPS-induced ERK activation and cytokine responses in macrophages [43]. IDO1 (indoleamine 2,3-dioxygenase 1) activity is the first and rate-limiting step in tryptophan catabolism in a variety of pathophysiological processes such as antimicrobial and antitumor defense, neuropathology, immunoregulation, and antioxidant activity [44]. The alpha M beta 2 integrin ITGAM (CD11b) is important in the adherence of neutrophils and monocytes to stimulated endothelium, and also in the phagocytosis of complement-coated particles. It has been shown that polymorphonuclear leukocytes play a pivotal role in early-phase immune response to bacterial or periprosthetic infection [45].
It is interesting to note that the antimicrobial activity may be beneficial for the treatment of acne vulgaris. Several essential oils have been used to treat acne. Proposed mechanisms for tea tree oil’s antiacne effects include antibacterial action against e.g. Propionibacterium acnes and anti-inflammatory properties [46]. Activities of ten essential oils towards P. acnes have been tested directly on the bacteria but not necessarily at the molecular level [47].
The cell surface molecule ABCC10 (ATP binding cassette subfamily C member 10) is a broad-acting transporter of xenobiotics, localized at the basolateral cell surface, as was shown with [3H]- docetaxel [48]. Finally, sulfotransferase enzymes, such as encoded by SULT1A2, catalyze the sulfate conjugation of many hormones, neurotransmitters, drugs, and xenobiotic compounds by increasing their solubility and facilitating their excretion [49].
The intriguing aspect of SO is that it shares the activities described in other medicinal plants, the active ingredients of curcumin, luteolin and aloe.
The genes SRC, CCND1, ODC1 and PCNA are genes that are equally activated by curcuminoids. Vogel and Pelletier (1815) [50] reported the isolation of a “yellow coloring-matter” from the rhizomes of Curcuma longa (turmeric) and named it curcumin. In turmeric, which is a member of the ginger family (Zingiberaceae), it is the principal curcuminoid, and chemically, curcumin is a diarylheptanoid. Recent research has shown that curcumin has a whole variety of medicinal activities: it modulates various signaling molecules, including inflammatory molecules, transcription factors, kinases, carrier proteins, cell survival proteins, drug resistance proteins, adhesion molecules, growth factors, receptors, cell-cycle regulatory proteins, chemokines, DNA, RNA, and metal ions [51].
Luteolin (3’,4’,5,7-tetrahydroxyflavone), a flavonoid extracted from Thymus vulgaris, inhibited PKCε and SRC kinase activity. In SKH-1 hairless mice, luteolin suppressed tumor incidence, multiplicity, and overall tumor size. Analysis of the skin by immunohistochemistry and immunoblotting showed that luteolin-treated groups had a substantial reduction in the levels of cyclooxygenase-2, tumor necrosis factor-alpha, and proliferating cell nuclear antigen compared with groups treated with only UVB. These data suggested that luteolin exerts potent chemopreventive activity against UVB-induced skin cancer mainly by targeting PKCε and SRC [52]. Luteolin reduced the activity of catalase and superoxide dismutase, and the level of oxidative damage, and lipid peroxidation, in lung tissue, and reduced LPS-induced activation of MAPK and NFκB pathways in mice [53]. Luteolin has also been demonstrated to inhibit lipopolysaccharide-induced tumor necrosis factor-α release and activation of the NFκB pathway in macrophages. In adipocytes, luteolin markedly enhanced PPARγ transcriptional activity in 3T3- L1 adipocytes, and luteolin-increased expression of adiponectin and leptin was blocked by GW9662, a PPARγ antagonist. The data suggest that luteolin influences insulin action and production of adipokines/cytokines in adipocytes by activating the PPARγ pathway [54]. Luteolin interfered with the regulation of estrogen signaling and cell cycle, consistent with the anti-estrogenic and anti-proliferative properties of luteolin in normal and malignant cells [55].
A third plant, aloe, was shown to reduce obesity-induced inflammation and the occurrence of metabolic disorders such as blood glucose and insulin resistance in obese mice. Obesity-induced inflammatory cytokine (IL-1b, IL-6, IL-12, TNFα) and chemokine (CX3CL1, CCL5) mRNA and protein were markedly decreased, as was macrophage infiltration and hepatic triglycerides by aloe. At the same time, aloe down-regulated the mRNA and protein of PPARγ/LXRα and 11β-HSD1 [56]. Treatment with aloe decreased the expression of these adipogenic genes and repressed adipocyte differentiation. In addition, aloe suppressed the differentiation of human mesenchymal stem cells into adipocytes by downregulating PPARγ and C/EBPα expressions indicating an anti-adipogenic effect [57].
Conclusion
This transcriptomic analysis shows the induction of four major activities, as determined by the main functions of induced genes and their correlations. These are 1) the retinoic acid pathway, 2) proliferation and cell cycle control, 3) cell growth and remodeling as well as 4) protection against microorganisms, antioxidant properties, and ultraviolet light. The pathways and gene correlations concern proliferation and differentiation and thus favor rejuvenating and anti-apoptotic activities, and cell renewal. The protective mechanisms induced by the treatment cover a broad range of targets: antimicrobial, antioxidant, anti-allergic and UVB protection. Some of these activities mimic features described with other plant extracts. The active ingredients of aloe, curcumin and the compound luteolin share properties that can be found in essential sandalwood oil, which may result in sandalwood sharing some of the beneficial activities.
This analysis provides the gene expression basis for a range of valuable properties, some of which are under clinical investigation or confirms traditional medicinal uses. The absence of adverse effects and allergic potential underlines the usefulness to combine traditional medicine with a scientific approach to achieve targeted applications in skin care.
Acknowledgments
The study 16E3411 was financed by SANTANOL and realized at GENEX in collaboration with the team of BIO-EC Laboratory and E. Lati. We are grateful for helpful discussions with A. Patatian at GENEX. No conflict of interest exists.
References
- Burdock GA, Carabin IG (2008) Safety assessment of sandalwood oil (Santalum album L.). Food Chem Toxicol 46: 421-432.
- Dwivedi C, Valluri HB, Guan X, Agarwal R (2006) Chemopreventive effects of alpha-santalol on ultraviolet B radiation-induced skin tumor development in SKH-1 hairless mice. Carcinogenesis 27: 1917-1922.
- Paulpandi M, Kannan S, Thangam R, Kaveri K, Gunasekaran P, et al. (2012) In vitro anti-viral effect of ß-santalol against influenza viral replication. Phytomedicine 19: 231-235.
- Sharma M, Levenson C, Bell RH, Anderson SA, Hudson JB, et al. (2014) Suppression of lipopolysaccharide-stimulated cytokine/chemokine production in skin cells by sandalwood oils and purified α-santalol and β-santalol. Phytother Res 28: 925-932.
- Palatty PL, Azmidah A, Rao S, Jayachander D, Thilakchand KR, et al. (2014) Topical application of a sandal wood oil and turmeric based cream prevents radiodermatitis in head and neck cancer patients undergoing external beam radiotherapy: a pilot study. Br J Radiol 87: 20130490.
- Kim J, Jang M, Shin E, Kim J, Lee SH, et al. (2016) Fumigant and contact toxicity of 22 wooden essential oils and their major components against Drosophila suzukii (Diptera: Drosophilidae). Pestic Biochem Physiol 133: 35-43.
- Fujisaki R, Kamei K, Yamamura M, Nishiya H, Inouye S, et al. (2012) In vitro and in vivo anti-plasmodial activity of essential oils, including hinokitiol. Southeast Asian J Trop Med Public Health 43: 270-279.
- Roh HS, Park KC, Park CG (2012) Repellent effect of santalol from sandalwood oil against Tetranychus urticae (Acari: Tetranychidae). J Econ Entomol 105: 379-385.
- Jayaprakasha GK, Rao LJM, Sakariah KK (2005) Chemistry and biological activities of C. longa. Trends Food Sci Tech 16: 533-548.
- Hussain AI, Anwar F, Hussain Sherazi ST, Przybylski R (2008) Chemical composition, antioxidant and antimicrobial activities of basil (Ocimum basilicum) essential oils depends on seasonal variations. Food Chemistry 108: 986-995.
- Sadler JJ, Smith JM, Zettler LW, Alborn HT, Richardson LW (2011) Fragrance composition of Dendrophylax lindenii (Orchidaceae) using a novel technique applied in situ. Euro J Environment Sci 1: 137-141
- Martins CdM, do Nascimento EA, de Morais SAL, de Oliveira A, Chang R, et al. (2015) Chemical constituents and evaluation of antimicrobial and cytotoxic activities of Kielmeyera coriacea. Mart. & Zucc. Essential Oils. Evidence-Based Complementary and Alternative Medicine Volume 2015: 842047.
- Moniodis J, Jones CG, Renton M, Plummer JA, Barbour EL et al. (2017) Sesquiterpene Variation in West Australian Sandalwood (Santalum spicatum). Molecules 22: 940-952.
- Vreeland AC, Levi L, Zhang W, Berry DC2 Noy N (2014) Cellular retinoic acid-binding protein 2 inhibits tumor growth by two distinct mechanisms. J Biol Chem 289: 34065-34073.
- Kim YW, Kim HJ, Bae SM, Kim YJ, Shin JC, et al. (2010) Time-course transcriptional profiling of human amniotic fluid-derived stem cells using microarray. Cancer Res Treat 42: 82-94.
- Si SP, Tsou HC, Lee X, Peacocke M (1995) Effect of cellular senescence and retinoic acid on the expression of cellular retinoic acid binding proteins in skin fibroblasts. Exp Cell Res 219: 243-248.
- Semple RK, Chatterjee VK, O'Rahilly S (2006) PPAR gamma and human metabolic disease. J Clin Invest 116: 581-589.
- Kang Y, Hengbo S, Jun L, Jun L, Wangsheng Z, et al. (2015) PPARG modulated lipid accumulation in dairy GMEC via regulation of ADRP gene. J Cell Biochem 116: 192-201.
- Chapellier B, Mark M, Messaddeq N, Calléja C, Warot X et al. (2002) Physiological and retinoid-induced proliferations of epidermis basal keratinocytes are differently controlled. EMBO J 21: 3402-3413.
- Elder JT, Aström A, Pettersson U, Tavakkol A, Griffiths CE et al. (1992) Differential regulation of retinoic acid receptors and binding proteins in human skin. J Invest Dermatol 98: 673-639.
- He G, Thuillier P, Fischer SM (2004) Troglitazone inhibits cyclin D1 expression and cell cycling independently of PPARgamma in normal mouse skin keratinocytes. J Invest Dermatol 123: 1110-1119.
- Baert B, De Spiegeleer B (2011) Local skin pharmacokinetics of talarozole, a new retinoic acid metabolism-blocking agent. Skin Pharmacol Physiol 24: 151-159.
- Bovenschen HJ, Otero ME, Langewouters AM, van Vlijmen-Willems IM, van Rens DWB et al. (2007) Oral retinoic acid metabolism blocking agent Rambazole for plaque psoriasis: an immunohistochemical study. J Dermatol 156:263-270.
- Wang X, Jiang L (2014) Effects of ornithine decarboxylase antizyme 1 on the proliferation and differentiation of human oral cancer cells. Int J Mol Med 34: 1606-1612.
- Jiang HL, Sun HF, Gao SP, Li LD, Hu X, et al. (2015) Loss of RAB1B promotes triple-negative breast cancer metastasis by activating TGF-ß/SMAD signaling. Oncotarget 6: 16352-16365.
- Wang J, Dong L, Xu L, Chu ES, Chen Y, et al. (2014) B cell CLL/lymphoma 6 member B inhibits hepatocellular carcinoma metastases in vitro and in mice. Cancer Lett 355: 192-200
- Tsai PJ, Davis J, Thompson K, Bryant-Greenwood G (2015) Visfatin/Nampt and SIRT1: Roles in Postterm Delivery in Pregnancies Associated With Obesity. Reprod Sci 22: 1028-1036.
- Kelsall IR, Voss M, Munro S, Cuthbertson DJ, Cohen PT (2011) R3F, a novel membrane-associated glycogen targeting subunit of protein phosphatase 1 regulates glycogen synthase in astrocytoma cells in response to glucose and extracellular signals. J Neurochem 118: 596-610.
- Kim JG, Kim EO, Jeong BR, Min YJ, Park JW, et al. (2010) Visfatin stimulates proliferation of MCF-7 human breast cancer cells. Mol Cells 30: 341-345.
- Ponomartsev NV, Enukashvily NE (2015) [The DDX5 protein is involved in cell proliferation and differentiation]. Tsitologiia 57: 111-118.
- Wang Y, Arribas-Layton M, Chen Y, Lykke-Andersen J, Sen GL (2015) DDX6 Orchestrates Mammalian Progenitor Function through the mRNA Degradation and Translational Pathway. Mol Cell 60: 118-130.
- Xu YF, Yang XQ, Lu XF, Guo S, Liu Y et al. (2014) Fibroblast growth factor receptor 4 promotes progression and correlates to poor prognosis in cholangiocarcinoma. Biochem Biophys Res Commun 446: 54-60.
- Kim BR, Dong SM, Seo SH, Lee JH, Lee JM, et al. (2014) Lysyl oxidase-like 2 (LOXL2) controls tumor-associated cell proliferation through the interaction with MARCKSL1. Cell Signal 26: 1765-1773.
- Pestov DG, Strezoska Z, Lau LF (2001) Evidence of p53-dependent cross-talk between ribosome biogenesis and the cell cycle: effects of nucleolar protein Bop1 on G(1)/S transition. Mol Cell Biol 21: 4246-4255.
- Carvalho SD, Chatterjee M, Coleman L, Clancy MA, Folta KM (2016) Analysis of Block of cell proliferation 1 (BOP1) activity in strawberry and Arabidopsis. Plant Sci 245: 84-93.
- Yin L, Xie Y, Yin S, Lv X, Zhang J et al. (2015) The S-nitrosylation status of PCNA localized in cytosol impacts the apoptotic pathway in a Parkinson's disease paradigm. PLoS One 12; 10: e0117546.
- Liu J, Ji X, Li Z, Zheng H, Zheng W (2015) Nestin overexpression promotes the embryonic development of heart and brain through the regulation of cell proliferation. Brain Res 1610: 1-11.
- Tiede S, Kloepper JE, Ernst N, Poeggeler B, Kruse C, et al. (2009) Nestin in human skin: exclusive expression in intramesenchymal skin compartments and regulation by leptin. J Invest Dermatol 129: 2711-2720.
- Makino Y, Tsuda M, Ohba Y, Nishihara H, et al. (2015) Tyr724 phosphorylation of ELMO1 by Src is involved in cell spreading and migration via Rac1 activation. Cell Commun Signal 13: 35.
- Kelwick R, Desanlis I, Wheeler GN, Edwards DR (2015) The ADAMTS (A Disintegrin and Metalloproteinase with Thrombospondin motifs) family. Genome Biol 16: 113.
- Storr SJ, Lee KW, Woolston CM, Safuan S, Green AR, et al. (2012) Calpain system protein expression in basal-like and triple-negative invasive breast cancer. Ann Oncol 23: 2289-2296.
- Polusani SR, Kalmykov EA, Chandrasekhar A, Zucker SN, Nicholson BJ (2016) Cell coupling mediated by connexin 26 selectively contributes to reduced adhesivity and increased migration. J Cell Sci 129: 4399-4410.
- Peng Q, O'Loughlin JL, Humphrey MB (2012) DOK3 negatively regulates LPS responses and endotoxin tolerance. PLoS One 7: e39967.
- Yeung AW, Terentis AC2, King NJ3, Thomas SR4 (2015) Role of indoleamine 2,3-dioxygenase in health and disease. Clin Sci (Lond) 129: 601-672.
- Kim GD, Lee SE, Yang H, Park HR, Son GW et al. (2014) ß2 integrins (CD11/18) are essential for the chemosensory adhesion and migration of polymorphonuclear leukocytes on bacterial cellulose. J Biomed Mater Res A 103: 1809-1817.
- Hammer KA (2015) Treatment of acne with tea tree oil (melaleuca) products: a review of efficacy, tolerability and potential modes of action. Int J Antimicrob Agents 45: 106-110.
- Zu Y, Yu H, Liang L, Fu Y, Efferth T, et al. (2010) Activities of ten essential oils towards Propionibacterium acnes and PC-3, A-549 and MCF-7 cancer cells. Molecules 15:3200-3210.
- Malofeeva EV, Domanitskaya N, Gudima M, Hopper-Borge EA (2012) Modulation of the ATPase and transport activities of broad-acting multidrug resistance factor ABCC10 (MRP7). Cancer Res 72:6457-6467.
- Fernández-Santander A, Gaibar M, Novillo A, Romero-Lorca A, Rubio M et al. (2013) Relationship between genotypes Sult1a2 and Cyp2d6 and tamoxifen metabolism in breast cancer patients. PLoS One 8: e70183.
- Vogel H, Pelletier J (1815) Curcumin –biological and medicinal properties. Journal de Pharmacie I: 289.
- Gupta SC, Patchva S, Koh W, Aggarwal BB (2012) Discovery of curcumin, a component of golden spice, and its miraculous biological activities. Clin Exp Pharmacol Physiol 39: 283-299.
- Byun S, Lee KW, Jung SK, Lee EJ, Hwang MK, et al. (2010) Luteolin inhibits protein kinase C(epsilon) and c-Src activities and UVB-induced skin cancer. Cancer Res 70: 2415-2423.
- Kuo MY, Liao MF, Chen FL, Li YC, Yang ML et al. (2011) Luteolin attenuates the pulmonary inflammatory response involves abilities of antioxidation and inhibition of MAPK and NF?B pathways in mice with endotoxin-induced acute lung injury. Food Chem Toxicol 49: 2660-2666.
- Ding L, Jin D, Chen X (2010) Luteolin enhances insulin sensitivity via activation of PPARγ transcriptional activity in adipocytes. J Nutr Biochem 21: 941-947.
- Markaverich BM, Shoulars K, Rodriguez MA (2011) Luteolin Regulation of Estrogen Signaling and Cell Cycle Pathway Genes in MCF-7 Human Breast Cancer Cells. Int J Biomed Sci 7: 101-111.
- Shin E, Shim KS, Kong H, Lee S, Shin S, et al. (2011) Dietary Aloe Improves Insulin Sensitivity via the Suppression of Obesity-induced Inflammation in Obese Mice. Immune Netw 11: 59-67.
- Subash-Babu P, Alshatwi AA (2012) Aloe-emodin inhibits adipocyte differentiation and maturation during in vitro human mesenchymal stem cell adipogenesis. J Biochem Mol Toxicol 26: 291-300.
 Spanish
Spanish  Chinese
Chinese  Russian
Russian  German
German  French
French  Japanese
Japanese  Portuguese
Portuguese  Hindi
Hindi