Research Article, Vector Biol J Vol: 3 Issue: 1
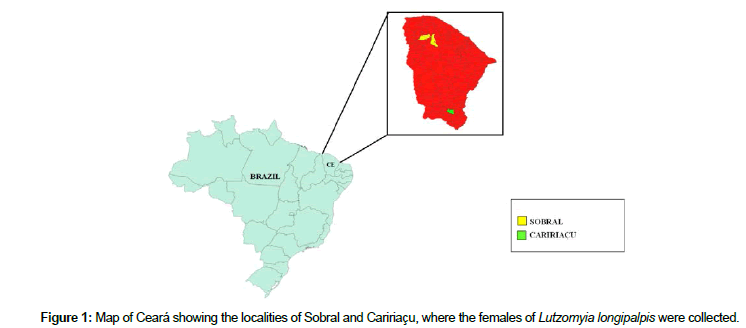
Genetic Structure of Sympatric Populations of Female Lutzomyia Longipalpis (Diptera: Psychodidae) In Sobral and Caririaçu, Ceará State, Brazil
Moises Thiago de Souza Freitas1*, César Raimundo Lima Costa Jr1, Lidiane Gomes da Silva1, Tereza Cristina Leal-Balbino2, Reginaldo Peçanha Brazil3 and Valdir de Queiroz Balbino1
1Department of Genética, Universidade Federal de Pernambuco, Av. Professor Moraes Rego S/N, Cidade Universitária, CEP: 50670-901 Recife, PE, Brasil
2Department of Microbiologia, Centro de Pesquisa Aggeu Magalhães, Av. Professor Moraes Rego S/N, Cidade Universitária, CEP: 50732-970 Recife, PE, Brasil
3Laboratório de Doenças Parasitarias, Instituto Oswaldo Cruz/Fiocruz, Av. Brasil, 4365 - Manguinhos, CEP: 21040-900 Rio de Janeiro, RJ, Brasil
*Corresponding Author : Moises Thiago de Souza Freitas
Departamento de Genética, Universidade Federal de Pernambuco, Av. Professor Moraes Rego S/N, Cidade Universitária, CEP: 50670-901 Recife, PE, Brasil
E-mail: moisesfrts@gmail.com
Received: December 05, 2017 Accepted: February 07, 2018 Published: February 27, 2018
Citation: Haruna A, Daskum AM (2018) Malaria and Haematological Parameters of Pregnant Women Attending General Hospital Geidam, Yobe State, Nigeria. Vector Biol J 3:1. doi: 10.4172/2473-4810.1000125
Abstract
Objectives: Accurate identification of these insects is critical to avoiding mistakes in the recognition of vector and non-vector species, so the invariant phenotypic patterns displayed by the females of L. longipalpis require the implementation of molecular methods of identification. Our goal was to examine genetic variation in the females of L. longipalpis using the SNPs reported in the males of this vector species.
Methods: We trapped sand flies near houses and domestic animal shelters at two localities in the state of Ceará, Brazil, using two to five CDC miniature light traps. Genomic DNA was separately extracted from each female using 100 μl of Chelex resin, and a 525 bp fragment of the period gene was used to assess genetic polymorphism in the two geographically isolated populations.
Results: The results indicate the presence of three fixed polymorphisms (T124C, C171T, and C424T) within this gene. Genetic structure analysis indicated that the studied L. longipalpis populations can be divided into two main subgroups (1S-like and 2S-like), with the ad hoc quantity supporting the number K = 2.
Conclusion: Collectively, our results suggest a direct relationship between the number of spots found in the males’ tergites and the genetic variation observed in the L. longipalpis females in the state of Ceará. The SNPs observed in the period gene will be useful for future studies of molecular eco-epidemiology in areas where these species occur in sympatry.
Keywords: Population genetics; Visceral leishmaniasis; sand fly; species complex; period gene; Northeast; Lutzomyia longipalpis
Introduction
The phlebotomine sand fly Lutzomyia longipalpis sensu lato is the principal vector of visceral leishmaniasis (VL) in the New World [1]. L. longipalpis males have either one pair of pale spots on the fourth tergite (the 1S morphotype) or two pairs on the third and fourth tergites (the 2S morphotype). The second pair on the third tergite may be smaller than the pair on the fourth and this is sometimes designated as an intermediate form [2]. There are reports of intermediate forms of L. longipalpis in some localities, especially around the northeast coast, indicating intraspecific polymorphism [2]. Several studies have shown that spot morphology cannot be used as a species-specific character [2,3]. Although these patterns of pale spots are not species-specific, they may be useful in identifying sympatric species in localities where intermediates are very rare, as occurs in Sobral [4], Jaiba, and Estrela de Alagoas [5]. The polymorphisms in the longer fragment of the period gene strongly suggest that this might also be the case in Bodocó and Caririaçu [6].
Molecular (microsatellite markers and speciation genes) and behavioral (sexual pheromones and courtship/mating sounds) analyses have also demonstrated the existence of differences between the 1S and 2S morphotypes and have provided further evidence of a L. longipalpis species complex [7]. Unlike the males, however, the females are morphologically undistinguishable: they do not present abdominal pale spots (either 1S or 2S), and this causes difficulties in studies of courtship and mating behaviors.
In Brazil, Ceará state has the third-highest number of VL cases, about 12.2% of the total reported in the country. Over the past 10 years, 4351 autochthonous cases were recorded in Ceará, and 578 of these cases (~13.3%) occurred in the municipality of Sobral [8]. In Sobral, two sympatric populations of L. longipalpis have exhibited differences in their copulatory courtship sounds (P3 and B), sex pheromones (9-methylgermacrene-B and cembrene), and ethological characters, as well as in their genetic variability as detected by molecular markers [6,7,9].
Multilocus analyses have also been performed on the L. longipalpis complex [10]. Among the studied loci, period (per) is widely known to be involved in the control of Drosophila circadian rhythms in eclosion and locomotor activity [11,12]. Also, period controls a feature of the “love song” that Drosophila males produce during courtship [13]. In sand flies, per has been utilized in population genetic studies to identify possible members of the L. longipalpis complex in Brazil [4,6,10].
Studies aiming to evaluate the genetic structure of the females of L. longipalpis populations have not been performed before, due to the females’ monomorphic character. Using analysis of microsatellite loci and pheromone types, however, identification of the females is now possible, and so the different population genetics of the females and males in this complex can now be studied [14]. In contrast to the females, several taxonomic, evolutionary, and behavioral studies have been performed using L. longipalpis males [4,15-18]. Accurate identification of these insects is critical to avoiding mistakes in the recognition of vector and non-vector species [19], so the invariant phenotypic patterns displayed by the females of L. longipalpis require the implementation of molecular methods of identification. Our goal was to examine genetic variation in the females of L. longipalpis using the SNPs reported in the males of this vector species [6].
Materials and Methods
Field collection and identification of phlebotomine sandflies
Sand fly trapping was carried out in Sobral (03°41′15″S; 40°21′5″W) and Caririaçu (07°02′31″S; 39°17′02″W) in the State of Ceará, Brazil. Both localities have a climate characterized as BSw′h′ according to the Köppen classification [20], with temperatures ranging from 23 °C to 36 °C and low annual rainfall (936 mm to 1100 mm). Both localities also are considered part of the Caatinga biome, with vegetation composed mainly of ligneous and herbaceous species with a high degree of xerophily.
Sand flies were trapped in the vicinity of houses and domestic animal shelters using two to five CDC miniature light traps positioned approximately 0.6 m from the ground. Collections were made from April 2013 to September 2014. Sand flies were identified according to Young and Duncan [21], and L. longipalpis females were separated for analysis. For the capture of the flies, consent from household owners was obtained before setting traps in and around their homes, and also near the shelters of domestic animals.
DNA extraction, PCR, and sequencing
Genomic DNA was separately extracted from each female using 100 μl of Chelex resin (Bio-Rad, Hercules, CA, USA) based on the protocol described by Costa Junior [6]. For each DNA sample, a 525 bp segment of the period gene [22] was amplified by PCR using a Master Mix Kit (Promega, Fitchburg, WI, USA). The amplicons were purified using the Genomic DNA Purification Wizard Kit (Promega, Fitchburg). Bi-directional sequencing reactions were performed on each purified PCR product using the BigDye terminator (Applied Biosystems, Foster City, CA, USA) and analyzed using an ABI3130 Sequence Analyzer (Applied Biosystems, Foster City). Each sample was sequenced in duplicate and the sequencing results evaluated with the Pregap4 program [23] based on the values of Phred 40 [24]. Heterozygous individuals were not observed. Consensus sequences were obtained using the Gap4 program [23] and deposited in GenBank under accession numbers KP144145 to KP144180.
Phylogenetic analysis
Of the 53 sequences of L. longipalpis females obtained for the period gene, 23 were from Sobral and 30 from Caririaçu. Sequence alignment was performed using the Muscle program present in MEGA v. 5.1 [25].
To generate a minimum evolution tree using MEGA v. 5.1 and to assess the consistency of the branches, the bootstrap test [26] with 1000 pseudo-replications was used. The sequences corresponding to the period gene in 1S and 2S male L. longipalpis from both localities (KF479047-KF479051, KF479106-KF479109, KF479075-KF479079, and KF479141-KF479145) were used as external groups.
The conserved and variables sites (parsimony-informative and singletons) were also verified using MEGA v. 5.1. For optimal viewing of polymorphisms, the parsimony-informative sites were exported into sequence logos using Weblogo v. 3.2 [27]. For the identification of SNPs, comparative analyses were done using the period sequences of L. longipalpis males with the 1S and 2S phenotypes, as described by Costa Junior [6].
Population structure
The genetic structuring of the L. longipalpis populations was verified using Structure v. 2.3 [28] with a model-based approach. Markov Chain Monte Carlo (MCMC) simulations were performed with 100,000 interactions of the burn-in period and followed by 1,000,000 steps. For each value of K (1 to 10), 10 interactions were performed to estimate the K values, and the most likely population (or cluster) number was determined by ΔK analysis, as described by Evanno [29].
Genetic diversity
Intra-population genetic diversity was assessed in terms of haplotype and nucleotide diversity, K value (number of genetic groups), number of polymorphic sites, and number of transitions and transversions; diversity was measured using DnaSP v. 4.0 [30] and Arlequin v. 3.5 [31].
Genetic differentiation was assessed with the pairwise fixation index Fst using Arlequin v. 3.5 [31]. The average number of substitutions per site among populations (Dxy), the total number of substitutions per site among populations (Da), the number of shared polymorphisms among populations (Ss), and the number of fixed differences among populations (Sf) were calculated using DnaSP v. 4.0 [30].
Results
Altogether, 53 L. longipalpis females were analyzed. The analyzed sequence region (525 bp) exhibited 63 (12%) polymorphic sites, including 39 (∼61.3%) parsimony-informative sites and 24 (∼38.7%) singletons. Among the polymorphic sites, 73% of the nucleotide substitutions were transitions and 27% were transversions (Table 1).
| Samples | N | Hd | π | NS | H | K | TS\TV | S | Pi |
|---|---|---|---|---|---|---|---|---|---|
| Sobral 1S | 10 | 1.0000 | 0.01636 | 27 | 10 | 8.55556 | 21\6 | 13 | 15 |
| Sobral 2S | 13 | 1.0000 | 0.01206 | 20 | 13 | 6.30769 | 16\5 | 7 | 14 |
| Caririaçu 1S | 12 | 1.0000 | 0.01721 | 28 | 12 | 9.0000 | 19\11 | 12 | 17 |
| Caririaçu 2S | 18 | 1.0000 | 0.01395 | 34 | 17 | 7.29412 | 26\11 | 19 | 16 |
| 1S-like full* | 22 | 1.0000 | 0.01692 | 37 | 22 | 8.84848 | 28\11 | 15 | 23 |
| 2S-like full* | 31 | 0.99785 | 0.01317 | 38 | 30 | 6.88602 | 29\12 | 20 | 19 |
| Total | 53 | 0.99927 | 0.02091 | 63 | 52 | 10.93759 | 46\17 | 24 | 39 |
π, nucleotide diversity; NS, number of polymorphic sites.
h, haplotype; K, average number of nucleotide difference; TS\TV, number of transitions\number of transversions.
S, singletons; Pi, parsimony-informative sites
*All females 1S-like and 2S-like.
Table 1: Intra-population genetic diversity measures for each sample.
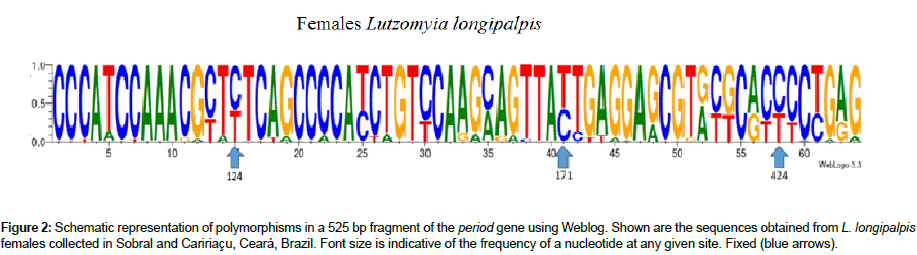
Three single nucleotide polymorphisms (SNPs) were identified within the 525 bp fragment of the period gene used in our analyses. Based on the SNPs, we found 22 1S-like specimens (12 FCAR and 10 FSOB) and 31 2S-like specimens (18 FCAR and 13 FSOB). All SNPs can be used to separate the phenotypes (1S and 2S). The SNPs identified at nucleotide positions T124C and C171T, the first and second nucleotides in the 1S and 2S phenotypes, respectively, are located within exon 1 and are fixed in L. longipalpis females from both localities in Ceará (Figure 1). The SNP identified at position C424T (within exon 2) is fixed in the L. longipalpis females from Sobral and Caririaçu, just as it is in L. longipalpis males.
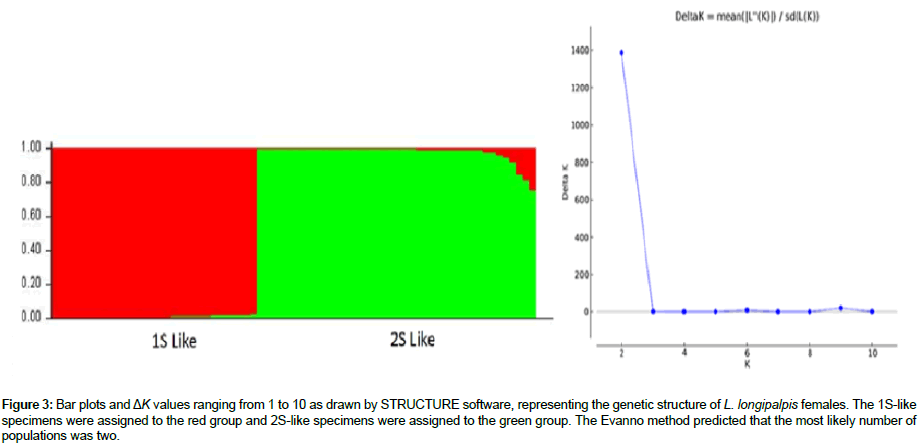
In the genetic structure analysis of L. longipalpis females, with each population assessed separately, two distinct genetic groups associated with abdominal spot patterns were observed, with each sequence possessing a probability (Q) greater than 80% of belonging to each genetic group (Figure 2). When sequences from both localities were combined, the genetic assignment test indicated the presence of two genetic groups associated with the 1S and 2S morphotypes, as suggested by the Δk peak that indicated the presence of two genetically distinct populations.
Figure 2: Schematic representation of polymorphisms in a 525 bp fragment of the period gene using Weblog. Shown are the sequences obtained from L. longipalpis females collected in Sobral and Caririaçu, Ceará, Brazil. Font size is indicative of the frequency of a nucleotide at any given site. Fixed (blue arrows).
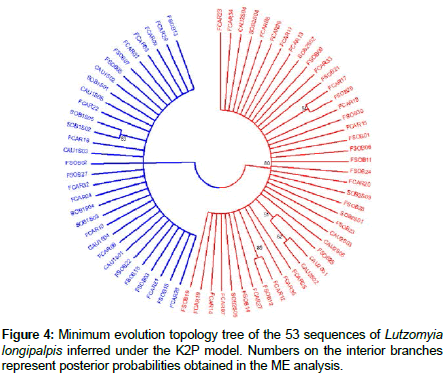
The minimum evolution analysis revealed two distinct and wellsupported clades with bootstrap values of 80%. This result indicates that the separation is associated with the abdominal spots (1S and 2S) of L. longipalpis males (Figures 3 and 4).
Figure 3: Bar plots and ΔK values ranging from 1 to 10 as drawn by STRUCTURE software, representing the genetic structure of L. longipalpis females. The 1S-like specimens were assigned to the red group and 2S-like specimens were assigned to the green group. The Evanno method predicted that the most likely number of populations was two.
Similarly, when Fst was used to verify the genetic structuring at the two localities (Caririaçu and Sobral), the presence of two morphotypes related to 1S and 2S was again observed. The L. longipalpis female 1S-like populations of Caririaçu and Sobral display low Fst values when compared to each other, forming a genetically similar group (Group 1S-like). A similar pattern was observed when comparing the 2S-like populations amongst themselves. However, the Fst values were the highest when comparing the two different phenotypes, even from the same locality (Table 2). Identical values of Fst were observed when populations of L. longipalpis males from the municipalities of Sobral and Caririaçu were examined [6].
| Populations | Fst | Dxy | Da | Ss | Sf | |
|---|---|---|---|---|---|---|
| Sobral 1S | Sobral 2S | 0.41906 | 0.02446 | 0.01025 | 8 | 1 |
| Caririaçu 1S | Caririaçu 2S | 0.48151 | 0.03004 | 0.01447 | 10 | 2 |
| Sobral 1S | Caririaçu 1S | 0.01095 | 0.01697 | 0.00019 | 18 | 0 |
| Sobral 1S | Caririaçu 2S | 0.40014 | 0.02526 | 0.01011 | 11 | 1 |
| Sobral 2S | Caririaçu 1S | 0.50063 | 0.02931 | 0.01467 | 8 | 3 |
| Sobral 2S | Caririaçu 2S | 0.00162 | 0.01302 | 0.00002 | 17 | 0 |
| 1S-like full* | 2S-like full* | 0.45395 | 0.02755 | 0.01251 | 14 | 1 |
Dxy: average number of nucleotide substitutions per site between populations.
Da: number of net nucleotide substitutions per site between populations.
Ss: number of shared polymorphisms between population pairs.
Sf: number of fixed differences between population pairs.
*All females 1S-like and 2S-like.
Table 2: Genetic differentiation among L. longipalpis populations.
Discussion
Analysis of the parsimony-informative sites indicated the presence of two group of female Lutzomyia longipalpis (1S-like and 2S-like). In a previous study, L. longipalpis males have been assessed for gene polymorphisms in the period gene with the aim of distinguishing the 1S and 2S morphotypes based on four SNPs in the populations of Sobral and Caririaçu [6]. These results indicate that there may be prezygotic barriers between the members of the L. longipalpis species complex, as suggested by Maingon [14] using microsatellite data and sex pheromones.
The fixed polymorphisms (T124C, C171T, and C424T) observed in the period gene will be particularly useful for the differentiation of 1S and 2S females in localities where they occur as sympatric species, just as they are in male L. longipalpis [6]. We also identified two sites with high frequencies that can be used in combination as markers for cryptic species occurring sympatrically in Sobral and Caririaçu. In male L. longipalpis, a fixed SNP causing a missense mutation in the amino acid sequence was previously identified in the gene paralytic, and it was able to separate the two main groups of L. longipalpis that produce different copulation songs [15,18]. The period gene is an effective tool to identify members of a species complex, as proved in Lutzomyia umbratilis [32].
The minimum evolution approach for L. longipalpis females collected in Caririaçu and Sobral indicated the presence of two genetically separated populations, and each collected sample belonged to either the 1S or the 2S morphotype. Our analyses also indicated that these two genetically distinct populations are correlated with abdominal spot patterns, as previously suggested for sympatric populations of L. longipalpis [6,10].
Conclusion
The SNPs observed in the period gene were useful to relate L. longipalpis females to the 1S and 2S phenotypes of the males, and they may be useful in the design of probes for future studies of molecular eco-epidemiology in areas where these species occur in sympatry.
Acknowledgements
This study was supported by Fundação de Amparo à Ciência e Tecnologia do Estado de Pernambuco (FACEPE) and Universidade Federal de Pernambuco.
References
- Grimaldi G, Tesh R, McMahon-Pratt D (1989) A review of the geographic distribution and epidemiology of leishmaniasis in the new world. Am J Trop Med Hyg 41: 687-725.
- Ward RD, Phillips A, Burnet B, Marcondes CB (1988) The Lutzomyia longipalpis complex: reproduction and distribution: Biosystematics of haematophagous insects. Systematics Association Special; Oxford, p. 257-269.
- Ward RD, Ribeiro AL, Ready PD, Murtagh A (1983) Reproductive isolation between different forms of Lutzomyia longipalpis (Lutz & Neiva) (Diptera: Psychodidae), the vector of Leishmania donovani chagasi Cunha & Chagas, and its significance to kala-azar distribution in South America. Mem Inst Oswaldo Cruz 78: 269-280.
- Bauzer LGSR, Souza NA, Ward RD, Kyriacou CP, Peixoto AA (2002) The period gene and genetic differentiation between three Brazilian populations of Lutzomyia longipalpis. Insect Mol Biol 11: 315-323.
- Araki AS, Vigoder FM, Bauzer LGSR, Ferreira GEM, Souza NA, et al. (2009) Molecular and behavioral differentiation among Brazilian populations of Lutzomyia longipalpis (Diptera: Psychodidae: Phlebotominae). Plos Negl Trop Dis 3: 1-12.
- Costa-Junior CRL, Freitas MTS, Figueiredo-Junior CAS, Aragao NC, Silva LG, et al. (2015) Genetic structuring and fixed polymorphisms in the gene period among natural populations of Lutzomyia longipalpis in Brazil. Parasit Vectors 8: 193.
- Bauzer LGSR, Souza NA, Maigon RDC, Peixoto AA (2007) Lutzomyia longipalpis in Brazil: a complex or a single species? A mini-review. Mem Inst Oswaldo Cruz 102: 1-12.
- Sistema de Informação de Agravos de Notificação (SINAN) (2016) Informações de Saúde (TABNET).
- Spiegel CN, Dias DBS, Araki AS, Hamilton JGC, Brazil RP, et al. (2016) The Lutzomyia longipalpis complex: a brief natural history of aggregation–sex pheromone communication. Parasit Vectors 9: 580.
- Araki AS, Ferreira GE, Mazzoni CJ, Souza NA, Machado RC, et al. (2013) Multilocus analysis of divergence and introgression in sympatric and allopatric sibling species of the Lutzomyia longipalpis complex in Brazil. Plos Negl Trop Dis 7: e2495.
- Hardin PE, Hall JC, Rosbash M (1990) Feedback of the Drosophila period gene product on circadian cycling of its messenger RNA levels. Nature 343: 536-340.
- Dunlap JC (1999) Molecular bases for circadian clocks. Cell 96: 271–290.
- Kyriacou CP, Hall JC (1980) Circadian rhythm mutations in Drosophila affect short-term fluctuations in the male’s courtship song. Proc Nat Acad Sci USA 77: 6929-6933.
- Maingon RD, Ward RD, Hamilton JG, Noyes HA, Souza N, et al. (2003) Genetic identification of two sibling species of Lutzomyia longipalpis (Diptera: Psychodidae) that produce distinct male sex pheromones in Sobral, Ceará State, Brazil. Mol Ecol 12: 1879-1894.
- Lins RMMA, Souza NA, Peixoto AA (2008) Genetic divergence between two sympatric species of the Lutzomyia longipalpis complex in the paralytic gene, a locus associated with insecticide resistance and lovesong production. 103: 736-740
- Rivas GB, Souza NA, Peixoto AA (2008) Analysis of the activity patterns of two sympatric sand fly siblings of the Lutzomyia longipalpis species complex from Brazil. Med Vet Entomol 22: 288-290.
- Silva MH, Nascimento MDSB, Leonardo FS, Rebêlo JMM, Pereira SRF (2011) Genetic differentiation in natural populations of Lutzomyia longipalpis (Lutz & Neiva) (Diptera: Psychodidae) with different phenotypic spot patterns on tergites in males. Neotrop Entomol 40: 501-506.
- Lins RMMA, Souza NA, Brazil RP, Maingon RDC, Peixoto AA (2012) Fixed differences in the paralytic gene define two lineages within the Lutzomyia longipalpis complex producing different types of courtship songs. PLoS One 7: 1-8.
- Marcondes CB (1998) Seria a taxonomia insignificante para a ecologia de vetores? Rev Soc Bras Med Trop 31: 492-493.
- Peel MC, Finlayson BL, McMahon TA (2007) Updated world map of the Koppen-Geiger climate classiï¬Âcation. Hydrol Earth Syst Sci11:1633-1644.
- Young DG, Duncan MA (1994) Guide to the identification and geographic distribution of Lutzomyia sand flies in Mexico, the West Indies, Central and South America (Diptera: Psychodidae). Mem Inst Oswaldo Cruz 54: 1-881.
- Mazzoni CJ, Gomes CA, Souza NA, De Queiroz RG, Justiniano SCB, et al. (2002) Molecular evolution of the period gene in sand flies. J Mol Evol 55: 553-562.
- Staden R (1996) The Staden sequence analysis package. Mol Biotechnol 5: 233-241
- Ewing B, Green P (1998) Base-calling of automated sequencer traces using Phred. II. Error probabilities. Genome Res 8: 186-194.
- Tamura K, Peterson D, Peterson N, Stecher G, Nei M, et al. (2011) MEGA 5: Molecular Evolutionary Genetics Analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol Biol Evol 28: 2731-2739.
- Felsenstein J (1985) Confidence limits on phylogenies: an approach using the bootstrap. Evolution 39: 783-791.
- Crooks GE, Hon G, Onchandia JM, Brenner SE (2004) WebLogo: a sequence logo generator. Genome Res 14: 1188-1190.
- Pritchard JK, Stephens P, Donnelly P (2000) Inference of population structure using multilocus genotype data. Genetics 155: 945-959.
- Evanno G, Regnaut S, Goudet, J (2005) Detecting the number of clusters of individuals using the software structure: a simulation study. Mol Ecol 14: 2611-2620.
- Rozas J, Sánches-DelBarrio JC, Messeguer X, Rozas R (2003) DnaSP, DNA polymorphism analyses by the coalescent and other methods. Bioinformatics 19: 2496-2497.
- Excoffier L, Lischer HEL (2010) Arlequin suite version 3.5: A new series of programs to perform population genetics analyses under Linux and Windows. Mol Ecol Resour 10: 564-567.
- De Souza Freitas MT, Ríos-Velasquez CM, Costa Junior CRL, Figueirêdo CAS, Aragão NC, et al. (2015) Phenotypic and genotypic variations among three allopatric populations of Lutzomyia umbratilis, main vector of Leishmania guyanensis. Parasit Vectors 8: 448.
 Spanish
Spanish  Chinese
Chinese  Russian
Russian  German
German  French
French  Japanese
Japanese  Portuguese
Portuguese  Hindi
Hindi